Welcome to the Recombinant Antibody Network
The Recombinant Antibody Network is a consortium of highly integrated technology centers at UCSF, the University of Chicago, and the University of Toronto, unified under a common goal to generate therapeutic grade recombinant antibodies at a proteome wide scale for biology and biomedicine.
Given that over half the human proteome is not annotated and that functional antibodies are not reliably available, a complete set of validated antibodies would greatly advance all areas of biology, including cancer therapy and infectious disease control. To undertake these challenges, RAN is systematically and comprehensively profiling families of protein targets using novel, modern high-throughput in vitro technology.
Latest Publications
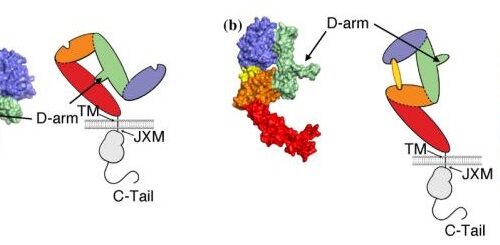
Adams J J; Mallette E; London M; Liang R J; van Dyk D; Pavlovic Z; Pot I; Geyer C R; Bruce H A; Blazer L L; Hokanson C A; Suits M D L; Singer A U; Sidhu S S
Dual targeting of inhibitory EGFR epitopes with synthetic antibodies in therapeutic-resistant cancers Journal Article
In: Protein Sci, vol. 35, no. 6, pp. e70645, 2026, ISSN: 1469-896X.
@article{pmid42178617,
title = {Dual targeting of inhibitory EGFR epitopes with synthetic antibodies in therapeutic-resistant cancers},
author = {Jarrett J Adams and Evan Mallette and Max London and Ryan J Liang and Dewald van Dyk and Zvezdan Pavlovic and Isabelle Pot and C Ronald Geyer and Heather A Bruce and Levi L Blazer and Craig A Hokanson and Michael D L Suits and Alexander U Singer and Sachdev S Sidhu},
doi = {10.1002/pro.70645},
issn = {1469-896X},
year = {2026},
date = {2026-06-01},
urldate = {2026-06-01},
journal = {Protein Sci},
volume = {35},
number = {6},
pages = {e70645},
abstract = {Therapeutic antibodies that inhibit the epidermal growth factor receptor (EGFR) are limited to a subset of EGFR-driven cancers. This is in part due to resistance mechanisms that attenuate efficacy. All approved therapeutic antibodies target the closed form of EGFR and compete with the ligand. However, tumors can be desensitized to these antibodies by upregulation of EGFR ligands or through EGFR mutations that uncouple kinase activity from ligand binding. We sought to overcome these resistance mechanisms by developing antibodies targeting alternative epitopes of EGFR. Using phage-displayed libraries, we developed two distinct antibodies, one that competed with EGF and another that did not. Crystal structures revealed that the competitive antibody bound to a site that overlapped the EGF-binding site, whereas the other antibody bound to the arm that induces receptor dimerization. Because the libraries used a common light chain, we were able to assemble a biparatopic antibody that incorporated both antigen-binding sites and thus targeted both epitopes on EGFR. We showed that the antibody that targeted the dimerization arm inhibited the growth of cancer cell lines that were resistant to the antibody that targeted the EGF-binding site. Moreover, the biparatopic antibody was more potent than the monoparatopic antibodies. Our results suggest that antibodies that target the dimerization arm of EGFR may be effective across a broader range of cancers than antibodies that target the EGF-binding site, and that a biparatopic antibody targeting both epitopes may be the most effective therapeutic for inhibiting aberrant EGFR signaling in cancer.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
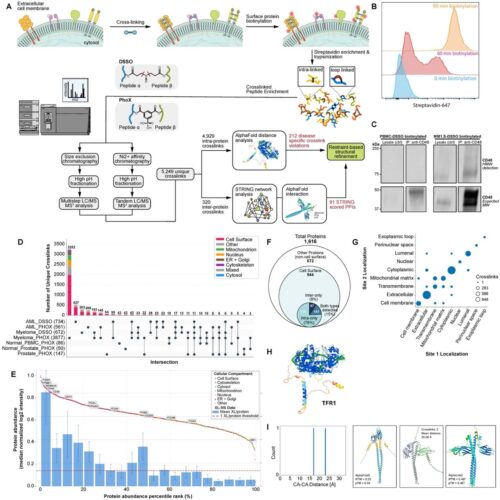
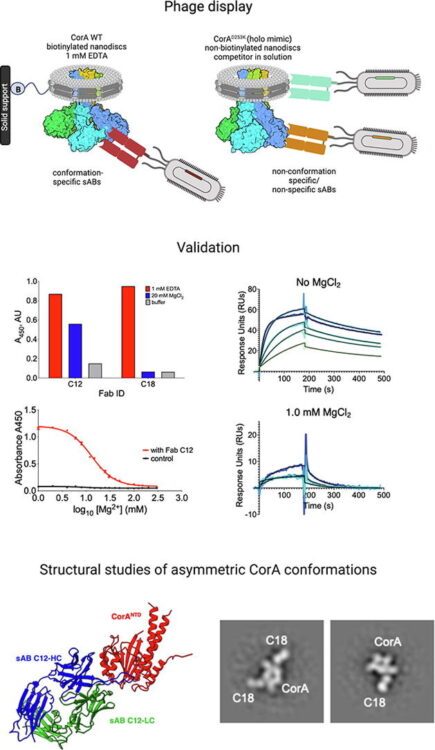
Kishishita A; Cismoski S; Grant T; Deo R; Prudhvi S; Sue C; Barpanda A; Yu C; Shenoy S; Berman S; Reeves A G; Li H; Liu T; Naik A; Biswas D; Jiao F; He Y; Hancock M; Dalal R; Zalevsky A; Hoopmann M R; Ye C J; Viner R I; Feng F; Mandal K; Moritz R L; Riesco I E; Sali A; Wells J A; Srivastava S; Huang L; Wiita A P
Extending structural surfaceomics to identify aberrant conformations of tumor surface proteins as potential immunotherapy targets Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid42239335,
title = {Extending structural surfaceomics to identify aberrant conformations of tumor surface proteins as potential immunotherapy targets},
author = {Audrey Kishishita and Sabine Cismoski and Tianna Grant and Rucha Deo and Sanjana Prudhvi and Catherine Sue and Abhilash Barpanda and Clinton Yu and Sanjyot Shenoy and Sarah Berman and Audrey G Reeves and Haolong Li and Tianyi Liu and Akul Naik and Deeptarup Biswas and Fenglong Jiao and Yi He and Matthew Hancock and Radhika Dalal and Arthur Zalevsky and Michael R Hoopmann and Chun Jimmie Ye and Rosa I Viner and Felix Feng and Kamal Mandal and Robert L Moritz and Ignacia Echeverria Riesco and Andrej Sali and James A Wells and Sanjeeva Srivastava and Lan Huang and Arun P Wiita},
doi = {10.64898/2026.05.15.721813},
issn = {2692-8205},
year = {2026},
date = {2026-05-01},
urldate = {2026-05-01},
journal = {bioRxiv},
abstract = {The complement of tumor cell surface proteins, or "surfaceome", is a rich source of potential immunotherapy targets. To move beyond expression-based target discovery, we previously described "structural surfaceomics," combining crosslinking mass spectrometry (XL-MS) with surface protein biotinylation to identify conformation-selective targets. In our prior work, we applied this method to a single model of acute myeloid leukemia (AML), identifying active integrin beta-2 as a promising target. Here, we expand structural surfaceomics to identify additional immunotherapy targets and surface protein biology across additional models of AML, multiple myeloma, and prostate cancer, as well as donor peripheral blood mononuclear cells. Utilizing these models and different chemical crosslinkers, we compile an extensive database of 5,209 crosslinks. We characterize both shared and unique crosslink-based features, identifying 1,612 disease model-specific crosslinks, including 212 potentially defining tumor-specific conformations based on distance constraint violations relative to AlphaFold predictions. We further implement a suite of emerging modeling tools to predict tumor-specific protein structures. We probe crosslinking patterns suggesting multiple myeloma-specific CD48 and AML-specific integrin α1/β4 heterodimer conformations. This work establishes a resource for cancer structural biology by implementation of structural surfaceomics. Our findings also point toward more realistic protein design models, potentially enabling systematic detection of targetable cancer-specific epitopes for next-generation immunotherapies.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
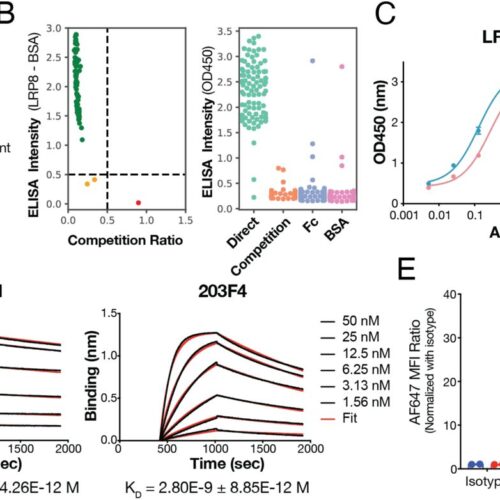
Zhao F; Inague A; Peters-Clarke T M; Chen Y; Ganjave S D; Zhang Y; Miao K; Yao Z; Wu Y; Seto M K C; Leung K K; Olzmann J A; Wells J A
Targeted extracellular degradation of LRP8 promotes ferroptosis in cancer cells Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid42239248,
title = {Targeted extracellular degradation of LRP8 promotes ferroptosis in cancer cells},
author = {Fangzhu Zhao and Alex Inague and Trenton M Peters-Clarke and Yifei Chen and Snehal D Ganjave and Yun Zhang and Kun Miao and Zi Yao and Yan Wu and Madison K C Seto and Kevin K Leung and James A Olzmann and James A Wells},
doi = {10.64898/2026.05.16.725645},
issn = {2692-8205},
year = {2026},
date = {2026-05-01},
urldate = {2026-05-01},
journal = {bioRxiv},
abstract = {Tumor reliance on antioxidant defenses creates a vulnerability to ferroptosis, yet strategies to therapeutically disable these systems remain limited. Here, we identify targeted degradation of the selenium uptake receptor LRP8 as an effective approach to decrease the abundance of the ferroptosis-protective enzyme glutathione peroxidase 4 (GPX4). Using bispecific cytokine receptor-targeting chimeras (KineTACs) that couple LRP8 to cytokine receptor internalization pathways, we selectively direct LRP8 to the lysosome for degradation. LRP8 degradation reduces the abundance of several selenoproteins, including GPX4, lowering the cellular threshold for lipid peroxidation and sensitizing cancer cells to ferroptosis. These findings establish receptor-mediated selenium uptake as a critical, targetable node in ferroptosis resistance and demonstrate that extracellular protein degradation can be leveraged to reprogram intracellular translational dependencies in cancer cells. More broadly, this work provides a framework for exploiting nutrient acquisition pathways to overcome therapy resistance.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Latest News
Recombinant Antibody Network Partners with Bristol Myers Squibb to Develop Novel Therapies
The Recombinant Antibody Network (RAN), a consortium comprising research groups from UC San Francisco, the…
Absolute Antibody Partners with the Recombinant Antibody Network to Facilitate Access to Engineered Recombinant Antibodies
Absolute Antibody Ltd., an industry-leading provider of recombinant antibody products and services, has announced a…
RAN to collaborate with Celgene on cancer therapeutics development
The RAN has recently signed a 3-year $25M agreement with the Celgene Corporation to develop next-generation,…
