Publications
View by year
- Publications: 2026
- Publications: 2025
- Publications: 2024
- Publications: 2023
- Publications: 2022
- Publications: 2021
- Publications: 2020
- Publications: 2019
- Publications: 2018
- Publications: 2017
- Publications: 2016
- Publications: 2015
- Publications: 2014
- Publications: 2013
- Publications: 2012
Search
Search for author, keywords, or any other term.
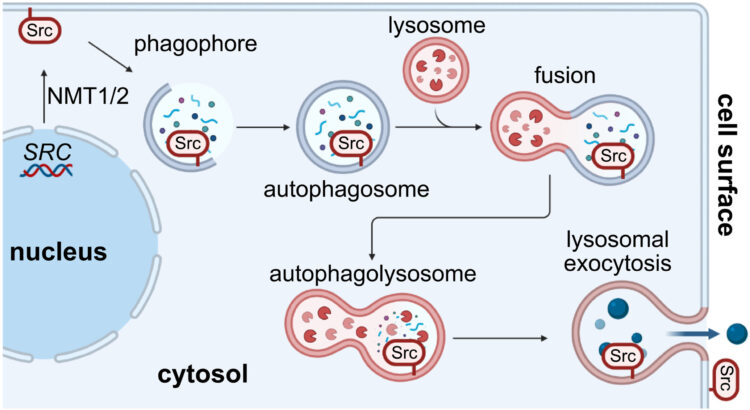
Delaveris, Corleone S; Loudermilk, Rita P; Pandey, Apurva; Remesh, Soumya G; Peters-Clarke, Trenton M; Ganjave, Snehal D; Dougherty, William J N; Delavan, Henry M; Wang, Chunyue; Ling, Jesse; Serrano, Juan Antonio Camara; Salangsang, Fernando; Ding, Chien-Kuang Cornelia; Greenland, Nancy; Chu, Carissa E; Porten, Sima; Steri, Veronica; Chou, Jonathan; Evans, Michael J; Leung, Kevin K; Wells, James A
Autophagolysosomal exocytosis inverts Src kinase onto the cell surface in cancer Journal Article
In: Science, vol. 391, no. 6790, pp. eaec1778, 2026, ISSN: 1095-9203.
@article{pmid41818370,
title = {Autophagolysosomal exocytosis inverts Src kinase onto the cell surface in cancer},
author = {Corleone S Delaveris and Rita P Loudermilk and Apurva Pandey and Soumya G Remesh and Trenton M Peters-Clarke and Snehal D Ganjave and William J N Dougherty and Henry M Delavan and Chunyue Wang and Jesse Ling and Juan Antonio Camara Serrano and Fernando Salangsang and Chien-Kuang Cornelia Ding and Nancy Greenland and Carissa E Chu and Sima Porten and Veronica Steri and Jonathan Chou and Michael J Evans and Kevin K Leung and James A Wells},
doi = {10.1126/science.aec1778},
issn = {1095-9203},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {Science},
volume = {391},
number = {6790},
pages = {eaec1778},
abstract = {Overexpression of the proto-oncogene Src is common to a wide variety of cancers. In this work, we found that Src is noncanonically translocated and inverted onto the cell surface in cancer, both in vitro and in vivo. We identified autophagolysosomal exocytosis (ALE) as a secretory mechanism prominent in cancer cell lines. Src represents the prototypical example of a family of membrane-anchored proteins that are transported by this process. Furthermore, this extracellular membrane-associated Src (eSrc) was found in primary tumors, and anti-Src antibody-based therapies mediated tumor cell killing in cell culture systems and in mouse xenograft models. Thus, intracellular -myristoylated proteins, prototypically Src, can be topologically inverted onto the cell surface in cancer and targeted with antibody therapeutics.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Alexander, J Andrew N; Chen, Shu-Yu; Mukherjee, Somnath; de Capitani, Mario; Irobalieva, Rossitza N; Rossi, Lorenzo; Agrawal, Parth; Kowal, Julia; Meirelles, Matheus A; Aebi, Markus; Reymond, Jean-Louis; Kossiakoff, Anthony A; Riniker, Sereina; Locher, Kaspar P
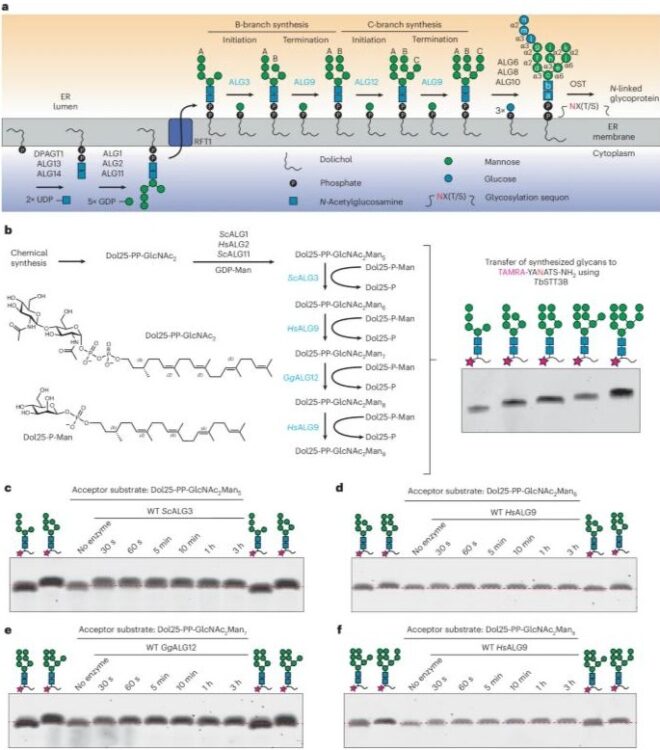
Structures of ALG3/9/12 reveal the assembly logic of the N-glycan oligomannose core Journal Article
In: Nat Chem Biol, 2026, ISSN: 1552-4469.
@article{pmid41807832,
title = {Structures of ALG3/9/12 reveal the assembly logic of the N-glycan oligomannose core},
author = {J Andrew N Alexander and Shu-Yu Chen and Somnath Mukherjee and Mario de Capitani and Rossitza N Irobalieva and Lorenzo Rossi and Parth Agrawal and Julia Kowal and Matheus A Meirelles and Markus Aebi and Jean-Louis Reymond and Anthony A Kossiakoff and Sereina Riniker and Kaspar P Locher},
doi = {10.1038/s41589-026-02164-7},
issn = {1552-4469},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {Nat Chem Biol},
abstract = {Asparagine-linked glycans are essential for the maturation and function of most eukaryotic secretory proteins. The biosynthesis and transfer of dolichylpyrophosphate-anchored GlcNAcManGlc glycan is a highly conserved process occurring in the endoplasmic reticulum (ER) membrane and involving over a dozen membrane proteins whose dysfunction is linked to congenital disorders of glycosylation (CDGs). Three membrane-integral mannosyltransferases, ALG3, ALG9 and ALG12, mediate four consecutive mannosylation reactions that convert GlcNAcMan to GlcNAcMan. Here, using chemoenzymatically synthesized lipid-linked glycan donor and acceptor analogs, we recapitulated this biosynthetic pathway in vitro. High-resolution cryo-electron microscopy structures of pseudo-Michaelis complexes of each step revealed how the branched glycan is accurately synthesized and unwanted side products are averted. Molecular dynamics simulations and mutagenesis studies uncovered a subtle but precise mechanism selecting the dolichylphosphomannose donor substrate over dolichylphosphoglucose, which is also present in the ER membrane. Our results also provide mechanistic explanations for enzyme dysfunction in CDGs and offer opportunities for N-glycan engineering.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Yao, Zi; Zhao, Fangzhu; Miao, Kun; Peters-Clarke, Trenton M; Zhang, Yun; Ganjave, Snehal D; Vázquez-Maldonado, Angel L; Wu, Yan; Kumru, Kaan; Jumaa, Hammam; Leung, Kevin K; Wells, James A
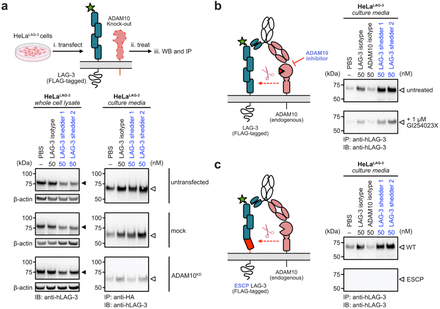
Targeted shedding of extracellular membrane proteins by induced protease recruitment Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid41756920,
title = {Targeted shedding of extracellular membrane proteins by induced protease recruitment},
author = {Zi Yao and Fangzhu Zhao and Kun Miao and Trenton M Peters-Clarke and Yun Zhang and Snehal D Ganjave and Angel L Vázquez-Maldonado and Yan Wu and Kaan Kumru and Hammam Jumaa and Kevin K Leung and James A Wells},
doi = {10.64898/2026.02.17.706468},
issn = {2692-8205},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {bioRxiv},
abstract = {Extracellular targeted protein degradation has emerged as a promising therapeutic modality to eliminate proteins of interest (POIs) at the cell surface, by using bifunctional molecules to recruit natural recycling receptors or membrane-bound E3 ligases that redirect POIs to the lysosome. Another natural mechanism involves extracellular proteases that cleave and shed extracellular domains. Here, we exploit this endogenous mechanism by engineering bispecific antibody , that recruit a classic sheddase ADAM10 to POIs, inducing selective ectodomain shedding. We first targeted the immune checkpoint receptor LAG-3 and observed robust depletion of surface LAG-3 accompanied by accumulation of soluble LAG-3 fragments in both engineered cell lines and primary human T cells. Using biochemical and imaging assays, we confirmed that this antibody-induced shedding is restricted to extracellular protease activity and occurs independently of lysosomal trafficking. Notably, induced shedding of LAG-3 on activated primary T cells partially alleviated inhibitory signaling and reinvigorated IFN secretion. We extended the scope of induced shedding by developing that recognize synthetic epitope-tags that enabling rapid assessment of substrate compatibility across diverse targets. Using this platform, we identified multiple immune modulatory cell-surface receptors, including IL6Rα, CD62L and MIC-A that can be targeted for shedding. In summary, this work establishes a new paradigm for targeted extracellular proteolysis and expands the toolkit for studying extracellular proteolysis with potential therapeutic benefit.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Erramilli, Satchal K; Nosol, Kamil; Pietrzak-Lichwa, Krzysztof; Schmandt, Nicolaus; Li, Tian; Tokarz, Piotr; Hou, Jingkai; Zhao, Minglei; Perozo, Eduardo; Kossiakoff, Anthony A
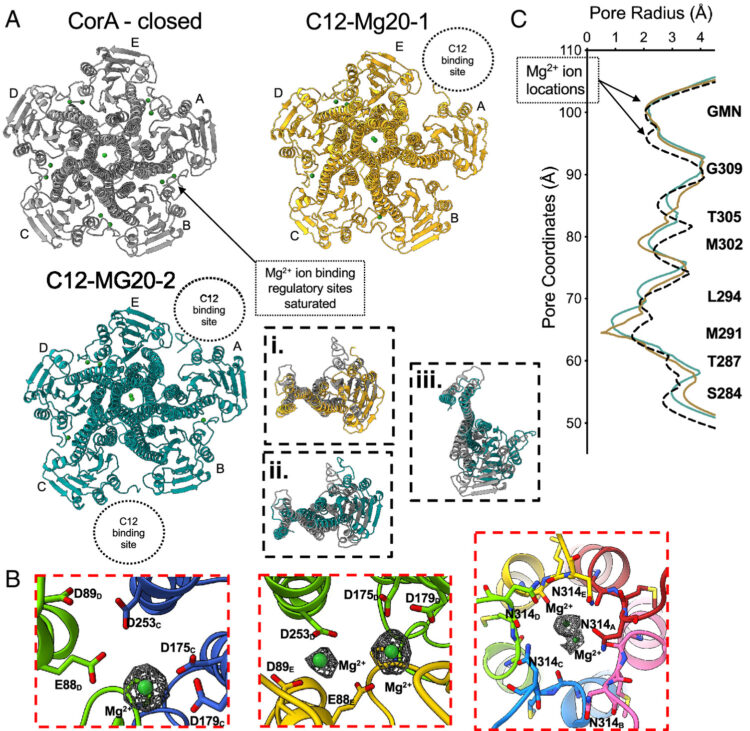
Conformational ensembles of the magnesium channel CorA reveal structural basis for channel gating Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 8, pp. e2512532123, 2026, ISSN: 1091-6490.
@article{pmid41701836,
title = {Conformational ensembles of the magnesium channel CorA reveal structural basis for channel gating},
author = {Satchal K Erramilli and Kamil Nosol and Krzysztof Pietrzak-Lichwa and Nicolaus Schmandt and Tian Li and Piotr Tokarz and Jingkai Hou and Minglei Zhao and Eduardo Perozo and Anthony A Kossiakoff},
doi = {10.1073/pnas.2512532123},
issn = {1091-6490},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {8},
pages = {e2512532123},
abstract = {In prokaryotes, CorA is the primary influx pathway for magnesium, a critical divalent cation in cellular physiology and biochemistry. Mechanistic studies show that homopentameric CorA is regulated through an intracellular [Mg]-dependent negative feedback loop, involving the asymmetric participation of individual subunits. To understand the connection between asymmetry and activation, we used single-particle cryo-EM to solve sixteen structures of nanodisc-reconstituted CorA. We utilized conformation-specific synthetic antibodies to stabilize subtle but significant conformational differences in the cryo-EM structures. Our results demonstrate that CorA exists as a set of conformational ensembles, where population size inversely correlates with intracellular Mg concentration. These ensembles include channels with a variety of pore conformations, both constricted and dilated, suggesting a spectrum of active CorA functional states. The ensembles connect asymmetric structural transitions in the cytoplasmic domain with conformational changes in the permeation pathway via an electrostatic network, ultimately controlling channel-gating events. We believe that these results establish a framework for understanding magnesium homeostasis in prokaryotic systems.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Adams, Jarrett J; Blazer, Levi L; Chung, Jacky; Karimi, Minoo; Davidson, Taylor; Blair, Bailey; Waddle, Carlos; Hokanson, Craig A; Bruce, Heather A; Singer, Alexander U; Tombak, Eva-Maria; Gildemann, Kiira; Tamberg, Nele; Kiiver, Kaja; Ustav, Mart; Ma, Yue; Colombo, Luigi; Huang, Lily Jun-Shen; Michnick, Stephen W; Moe, Orson W; Sidhu, Sachdev S
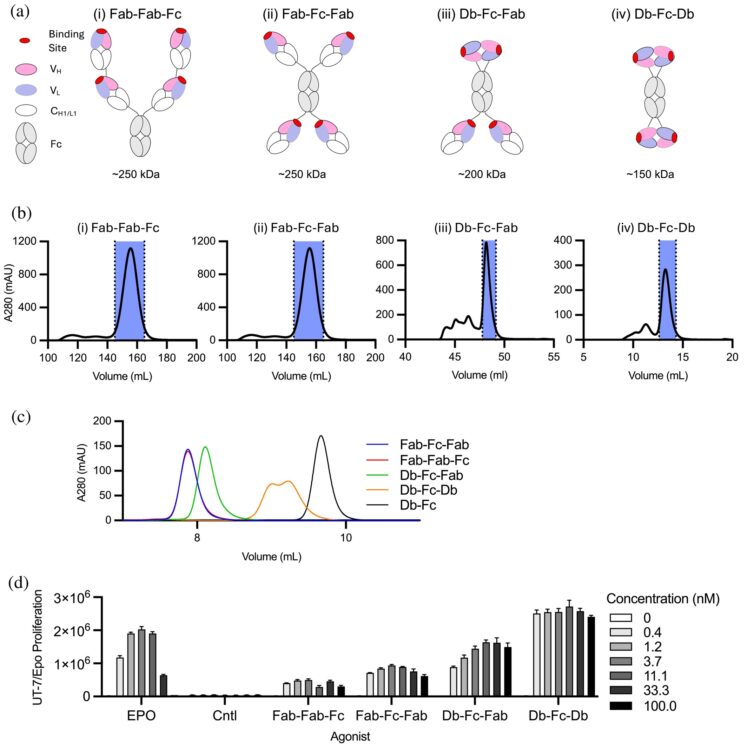
Tetravalent antibodies are more potent and efficacious erythropoiesis-stimulating agents than erythropoietin in vivo Journal Article
In: Protein Sci, vol. 35, no. 2, pp. e70462, 2026, ISSN: 1469-896X.
@article{pmid41556618,
title = {Tetravalent antibodies are more potent and efficacious erythropoiesis-stimulating agents than erythropoietin in vivo},
author = {Jarrett J Adams and Levi L Blazer and Jacky Chung and Minoo Karimi and Taylor Davidson and Bailey Blair and Carlos Waddle and Craig A Hokanson and Heather A Bruce and Alexander U Singer and Eva-Maria Tombak and Kiira Gildemann and Nele Tamberg and Kaja Kiiver and Mart Ustav and Yue Ma and Luigi Colombo and Lily Jun-Shen Huang and Stephen W Michnick and Orson W Moe and Sachdev S Sidhu},
doi = {10.1002/pro.70462},
issn = {1469-896X},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {Protein Sci},
volume = {35},
number = {2},
pages = {e70462},
abstract = {Recent studies have shown that tetravalent antibodies are potent and efficacious agonists of the erythropoietin (EPO) receptor (EPOR) both in vitro and in vivo. To identify antibody-based erythropoiesis-stimulating agents (ESAs) with therapeutic potential, we evaluated various tetravalent antibody formats for EPOR agonism and key biophysical properties necessary for biologic drug development. We identified two distinct tetravalent antibody formats that strongly stimulated the growth of UT7/Epo cells, which rely on EPOR signaling for proliferation. Moreover, one of these formats exhibited ideal biophysical characteristics for drug development. This format consisted of a diabody (Db) and two antigen-binding fragment (Fab) arms fused to the N- and C-termini of an Fc domain, respectively, to form a tetravalent Db-Fc-Fab (EPRA-0322). In a mouse model expressing the human EPOR, EPRA-0322 induced erythropoiesis with greater potency, efficacy, and duration than darbepoetin, a hyperglycosylated EPO currently used in clinical practice. These findings highlight tetravalent antibodies, and the Db-Fc-Fab format in particular, as promising next-generation ESAs suitable for large-scale production and clinical use.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Lin, Zhi; Ngo, Wayne; Chou, Yu-Ting; Wu, Harry; Susa, Katherine J; Jun, Young-Wook; Bivona, Trever G; Doudna, Jennifer A; Wells, James A
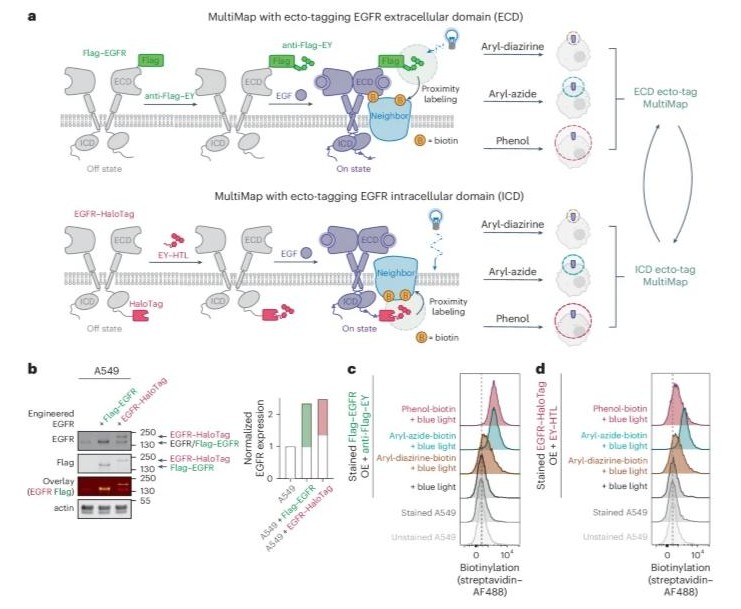
Temporal photoproximity labeling of ligand-activated EGFR neighborhoods using MultiMap Journal Article
In: Nat Chem Biol, vol. 22, no. 2, pp. 192–204, 2026, ISSN: 1552-4469.
@article{pmid41254216,
title = {Temporal photoproximity labeling of ligand-activated EGFR neighborhoods using MultiMap},
author = {Zhi Lin and Wayne Ngo and Yu-Ting Chou and Harry Wu and Katherine J Susa and Young-Wook Jun and Trever G Bivona and Jennifer A Doudna and James A Wells},
doi = {10.1038/s41589-025-02076-y},
issn = {1552-4469},
year = {2026},
date = {2026-02-01},
urldate = {2025-11-01},
journal = {Nat Chem Biol},
volume = {22},
number = {2},
pages = {192--204},
abstract = {Photoproximity labeling proteomics (PLP) methods have recently shown that cell surface receptors can form lateral interactome networks. Here, we present a paired set of PLP workflows that dynamically track neighborhood changes for oncogenic epidermal growth factor receptor (EGFR) over time, both outside and inside of cells. We achieved this by augmenting the multiscale PLP workflow we call MultiMap, where three photoprobes with different labeling ranges were photoactivated by one photocatalyst, eosin Y, anchored extracellularly and intracellularly on EGFR. We identified hundreds of neighboring proteins that changed within minutes to over 1 h after the addition of EGF. These neighborhoods reveal dynamic interactomes during early, middle and late signaling that drive phosphorylation, internalization, degradation and transcriptional regulation. This rapid 'molecular photographic' labeling approach provides snapshots of signaling neighborhoods, revealing their dynamic nature and potential for drug targeting.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Adams, Jarrett J.; Blazer, Levi L.; Chen, Gang; Chung, Jacky; Mallette, Evan; Singer, Alexander U.; Mendiratta, Sanjeev K.; Bandyopadhyay, Sanjay; Singh, Arun K.; Kalita, Pankaj; Karimi, Minoo; Cardarelli, Lia; Geyer, C. Ronald; Michnick, Stephen W.; Sidhu, Sachdev S.
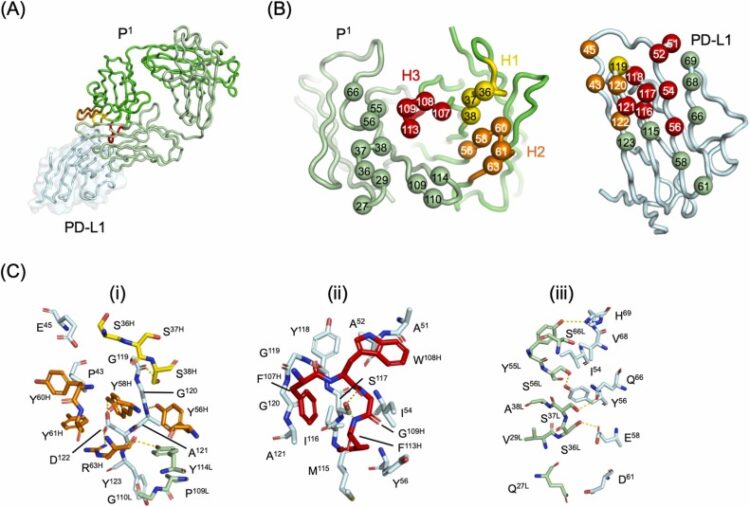
A tetravalent PD-L1/VEGF bispecific antibody blocks tumor growth In Vivo Journal Article
In: Disease and Therapeutics, vol. 2, 2026, ISSN: 3050-9998.
@article{Adams2026,
title = {A tetravalent PD-L1/VEGF bispecific antibody blocks tumor growth In Vivo},
author = {Jarrett J. Adams and Levi L. Blazer and Gang Chen and Jacky Chung and Evan Mallette and Alexander U. Singer and Sanjeev K. Mendiratta and Sanjay Bandyopadhyay and Arun K. Singh and Pankaj Kalita and Minoo Karimi and Lia Cardarelli and C. Ronald Geyer and Stephen W. Michnick and Sachdev S. Sidhu},
doi = {10.1016/j.dist.2026.100006},
issn = {3050-9998},
year = {2026},
date = {2026-01-06},
urldate = {2026-00-00},
journal = {Disease and Therapeutics},
volume = {2},
publisher = {Elsevier BV},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Montaño, José L; Tejus, Vishnu R; Xu, Vee; Condon, Andrew C; Ecker, Andrew K; Langen, Andreas; Leung, Kevin K; Zaro, Balyn W
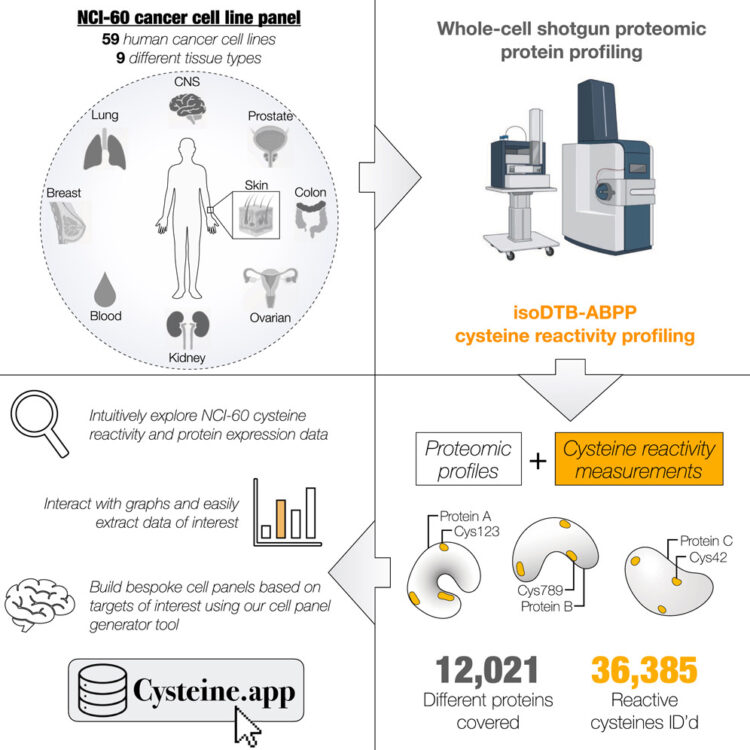
An interactive resource mapping the proteome and reactive cysteine landscape across the NCI-60 reveals cell and tissue-specific profiles Journal Article
In: Cell Chem Biol, vol. 33, no. 1, pp. 117–131.e5, 2026, ISSN: 2451-9448.
@article{pmid41478280,
title = {An interactive resource mapping the proteome and reactive cysteine landscape across the NCI-60 reveals cell and tissue-specific profiles},
author = {José L Montaño and Vishnu R Tejus and Vee Xu and Andrew C Condon and Andrew K Ecker and Andreas Langen and Kevin K Leung and Balyn W Zaro},
doi = {10.1016/j.chembiol.2025.12.003},
issn = {2451-9448},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Cell Chem Biol},
volume = {33},
number = {1},
pages = {117--131.e5},
abstract = {The NCI-60 cancer cell line panel is one of the most extensively characterized and publicly accessible resources in cancer research. Here, we expand this platform by generating a comprehensive proteomic and cysteine reactivity atlas using shotgun proteomics and quantitative chemoproteomics. We quantified over 12,000 proteins and identified more than 36,000 reactive cysteines, including over 10,000 hyperreactive sites, across the panel. Our analyses reveal widespread heterogeneity in cysteine reactivity, while conserved hyperreactive cysteines strongly correlate with functional roles. Tissue-specific cysteine reactivity occurred independently of protein abundance, highlighting context-dependent regulation. To enable exploration and integration with existing datasets, we developed an interactive online database that allows users to visualize protein and cysteine coverage and design custom cell line panels. Together, these data and tools illuminate the landscape of cysteine reactivity in cancer and provide a foundational resource to advance future studies in protein function, redox biology, and covalent drug discovery.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Kumru, Kaan; Yao, Zi; Holmes, Brandon B; Zhao, Fangzhu; Zhang, Yun; Ferrara, Emilio; Peters-Clarke, Trenton M; Leung, Kevin K; Wells, James A
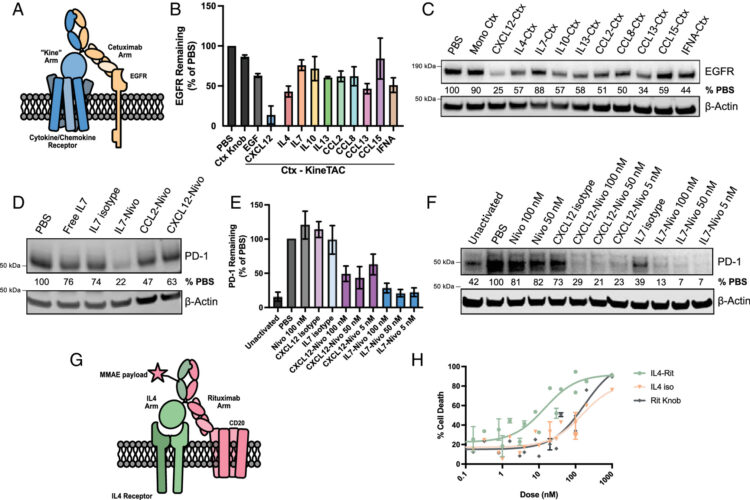
A cytokine receptor-targeting chimera toolbox for expanding extracellular targeted protein degradation Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 4, pp. e2524129123, 2026, ISSN: 1091-6490.
@article{pmid41564137,
title = {A cytokine receptor-targeting chimera toolbox for expanding extracellular targeted protein degradation},
author = {Kaan Kumru and Zi Yao and Brandon B Holmes and Fangzhu Zhao and Yun Zhang and Emilio Ferrara and Trenton M Peters-Clarke and Kevin K Leung and James A Wells},
doi = {10.1073/pnas.2524129123},
issn = {1091-6490},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {4},
pages = {e2524129123},
abstract = {Extracellular targeted protein degradation (eTPD) is an important new modality for manipulating the extracellular proteome. However, most eTPD receptors are expressed broadly or are restricted to the liver, limiting specific degradation in other tissues. Cytokine receptor-targeting chimeras (kineTACs) are genetically encoded bispecifics for eTPD that fuse a natural ligand like CXCL12 to an antibody, directing soluble or membrane proteins for lysosomal degradation using the widely expressed chemokine receptor CXCR7 (K. Pance , , 273-281 (2023)]. Here, we dramatically expand the kineTAC toolbox by constructing 81 different kineTACs based on an unbiased list of cytokines, chemokines, and growth factors. Remarkably, 55 of these expressed at suitable levels for analysis without any optimization. Many of these kineTACs bind receptors that have unique cell-type expression profiles, allowing for eTPD in specific cells and tissues, and some were more potent than the original CXCL12-based kineTAC against specific targets. We further show the internalizing capability of a kineTAC can enhance the performance of antibody drug conjugates. We believe these simple, genetically encoded tools will be useful for expanding the applications for optimized or cell type-selective eTPD.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
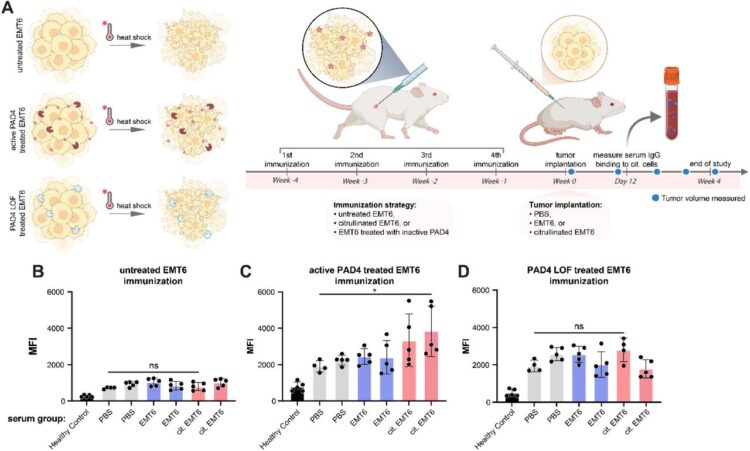
Kong, Sophie; Peters-Clarke, Trenton M; Delaveris, Corleone S; Phojanakong, Paul; Steri, Veronica; Wells, James A
Cellular consequences, citrullination substrates, and antigenicity resulting from wild-type and targeted PAD4 on cell surfaces Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid41542531,
title = {Cellular consequences, citrullination substrates, and antigenicity resulting from wild-type and targeted PAD4 on cell surfaces},
author = {Sophie Kong and Trenton M Peters-Clarke and Corleone S Delaveris and Paul Phojanakong and Veronica Steri and James A Wells},
doi = {10.64898/2026.01.05.696859},
issn = {2692-8205},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {bioRxiv},
abstract = {Protein arginine deiminase-4 (PAD4) catalyzes hydrolysis of arginine to citrulline in proteins that promotes widespread changes in cellular phenotypes through transcriptional regulation that can induce innate immunity and promote cancer. Overexpression and hyperactivity of PAD4 leads to a form of cell death called NETosis that releases PAD4 to the extracellular space. In excess, release of PAD4 is believed to be a major cause of various autoimmune diseases through the generation of anti-citrulline protein antibodies (ACPAs). Little is known about the specific protein substrates that become citrullinated and lead to autoimmunity, but there is growing evidence that PAD4 can be localized to the cell surface in response to inflammation. Here, we further characterize the cellular consequences for exogenous treatment with PAD4 showing that it induces morphological changes that increase cell migration, a hallmark of cancer. We then devised a more simplified and robust proteomics approach to identify PAD4 substrates. We identified some 1000 endogenously citrullinated peptides from 500 proteins, and 3000 citrullinated peptides from 1300 proteins upon exogenous addition of PAD4 both inside and outside of cells. This extracellular set can be further augmented by targeting PAD4 to a cancer target, HER2, using a binding protein conjugate. Finally, we studied how citrullinated cells can induce a robust humoral response in a syngeneic vaccine model to produce ACPAs. We believe these studies further our understanding of cell phenotypic consequences of extracellular PAD4 and new PAD4 substrates both inside and outside of cells that are potential neoepitopes for generation of ACPAs.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
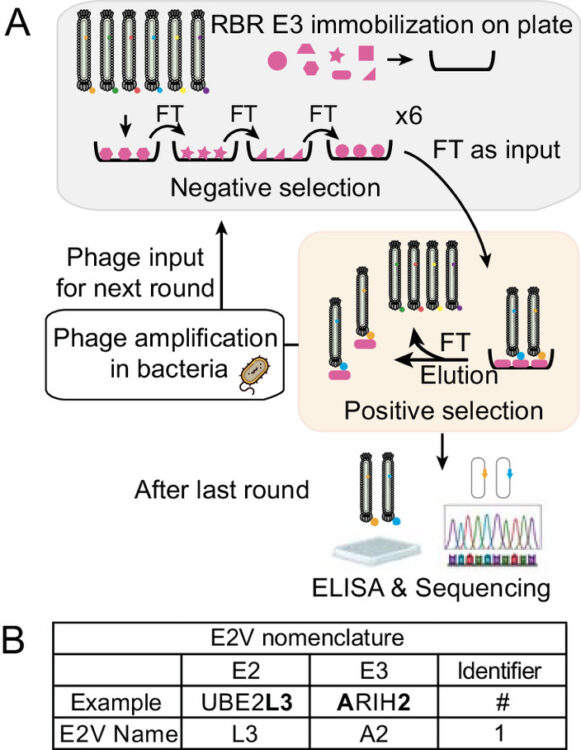
Du, Jiale; Andree, Gisele A; Horn-Ghetko, Daniel; Stier, Luca; Singh, Jaspal; Kostrhon, Sebastian; Kiss, Leo; Mann, Matthias; Sidhu, Sachdev S; Schulman, Brenda A
E2 variants for probing E3 ubiquitin ligase activities Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 1, pp. e2524899122, 2026, ISSN: 1091-6490.
@article{pmid41481455,
title = {E2 variants for probing E3 ubiquitin ligase activities},
author = {Jiale Du and Gisele A Andree and Daniel Horn-Ghetko and Luca Stier and Jaspal Singh and Sebastian Kostrhon and Leo Kiss and Matthias Mann and Sachdev S Sidhu and Brenda A Schulman},
doi = {10.1073/pnas.2524899122},
issn = {1091-6490},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {1},
pages = {e2524899122},
abstract = {E3 ligases partner with E2 enzymes to regulate vast eukaryotic biology. The hierarchical nature of these pairings, with >600 E3s and ~40 E2s in humans, necessitates that E2s cofunction with numerous different E3s. Here, focusing on E3s in the RING-between-RING (RBR) family and their partner UBE2L3 and UBE2D-family E2s, we report an approach to interrogate selected pathways. We screened phage-displayed libraries of structure-based E2 variants (E2Vs) to discover enzymes with enhanced affinity and specificity toward half of all RBR E3 ligases (ARIH1, ARIH2, ANKIB1, CUL9, HOIL1, HOIP, and RNF14). Collectively, these E2Vs allowed distinguishing actions of different cofunctioning E3s, obtaining high-resolution cryogenic Electron Microscopy (cryo-EM) structures of an RBR E3 in the context of a substrate-bound multiprotein complex, and profiling an endogenous RBR E3 response to an extracellular stimulus. Overall, we anticipate that E2V technology will be a generalizable tool to enable in-depth mechanistic and structural analysis of E3 ligase functions, and mapping their activity states and protein partners in cellular signaling cascades.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Han, Chengxiao; Weng, Yicheng; Zheng, Qingyun; Qu, Qian; Erramilli, Satchal K; Su, Zhen; Duan, Yujuan; Han, Yunxi; Zhai, Xiaoguo; Li, Jingxian; Kossiakoff, Anthony A; Pan, Man; Zhao, Minglei; Liu, Lei; Yu, Yuanyuan
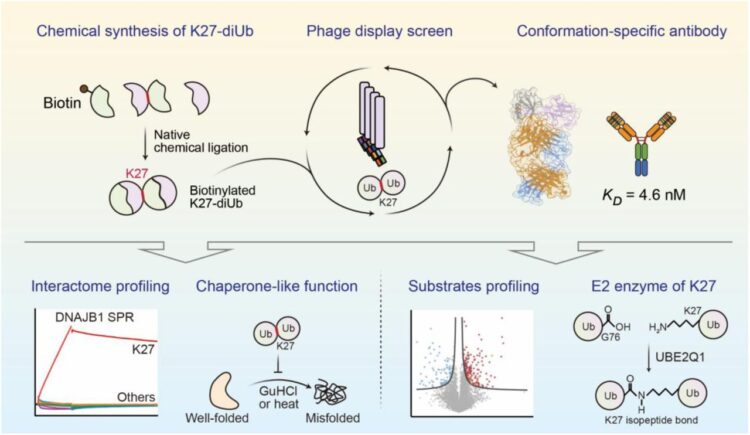
Conformation-specific Antibody Deciphers K27-linked Ubiquitination in Chaperone-Mediated Proteostasis Journal Article
In: bioRxiv, 2025, ISSN: 2692-8205.
@article{pmid41446272,
title = {Conformation-specific Antibody Deciphers K27-linked Ubiquitination in Chaperone-Mediated Proteostasis},
author = {Chengxiao Han and Yicheng Weng and Qingyun Zheng and Qian Qu and Satchal K Erramilli and Zhen Su and Yujuan Duan and Yunxi Han and Xiaoguo Zhai and Jingxian Li and Anthony A Kossiakoff and Man Pan and Minglei Zhao and Lei Liu and Yuanyuan Yu},
doi = {10.64898/2025.12.18.695067},
issn = {2692-8205},
year = {2025},
date = {2025-12-01},
urldate = {2025-12-01},
journal = {bioRxiv},
abstract = {Lysine 27 (K27)-linked polyubiquitination plays critical yet incompletely defined roles in proteostasis, innate immunity, and disease progression; however, investigations into this process have long been hindered by its extremely low abundance and the lack of conformation-specific enrichment tools. Herein, we describe the development of a long-sought conformation-specific antibody, K27-IgG, which can selectively recognize-among all ubiquitin chain types-the unique architecture of K27-linked polyubiquitin (K27-polyUb) characterized by a distinct buried K27-isopeptide bond, with high affinity (KD = 4.66 nM). This antibody was derived from synthetic antibodies initially generated via phage display, using chemically synthesized K27-linked diubiquitin (K27-diUb) as the antigen. High-resolution co-crystal structures uncovered the unique K27-diUb interface targeted by these sAbs. Subsequent reformatting of these sAbs into a full-length human immunoglobulin G (IgG) scaffold yielded K27-IgG, notably exhibiting markedly enhanced affinity without compromising selectivity. Using K27-IgG as a tool, we achieved sensitive detection and immunoprecipitation (IP) of endogenous K27-polyUb in cells, and delineated the intracellular interaction landscape of K27-polyUb through complementary proteomic approaches. Two key findings emerged: 1) The molecular chaperone DNAJB1 is a specific reader of K27-linked ubiquitin chains (but not other linkages) and that K27-polyUb chains themselves exhibit chaperone-like activity, suggesting a novel mechanism by which K27-polyUb regulates chaperone-mediated proteostasis; 2) The E2 enzyme UBE2Q1 assembles K27-diUb, identifying it as a potential writer for this ubiquitin chain topology. Collectively, this study establishes K27-IgG as a robust tool for deciphering the K27-linked ubiquitin code, thereby opening new avenues for investigating the biological functions of K27-linked polyubiquitination.nnHIGHLIGHTS: First K27-linkage conformation-specific antibody with nanomolar affinity overcomes a major barrier in the field.K27-IgG unlocks functional mapping of the K27 ubiquitin landscape under proteotoxic stress.Molecular chaperone DNAJB1 is a selective "reader" of K27-linked ubiquitin chains.K27 chains possess intrinsic chaperone activity, enabling protein refolding and suppressing aggregation.E2 enzyme UBE2Q1 is a "writer" that directly assembles K27-linked ubiquitin chains.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Zinkle, Allen P; Batista, Mariana Bunoro; Herrera, Carmen M; Erramilli, Satchal K; Kloss, Brian; Ashraf, Khuram U; Nosol, Kamil; Zhang, Guozhi; Cater, Rosemary J; Marty, Michael T; Kossiakoff, Anthony A; Trent, M Stephen; Nygaard, Rie; Stansfeld, Phillip J; Mancia, Filippo
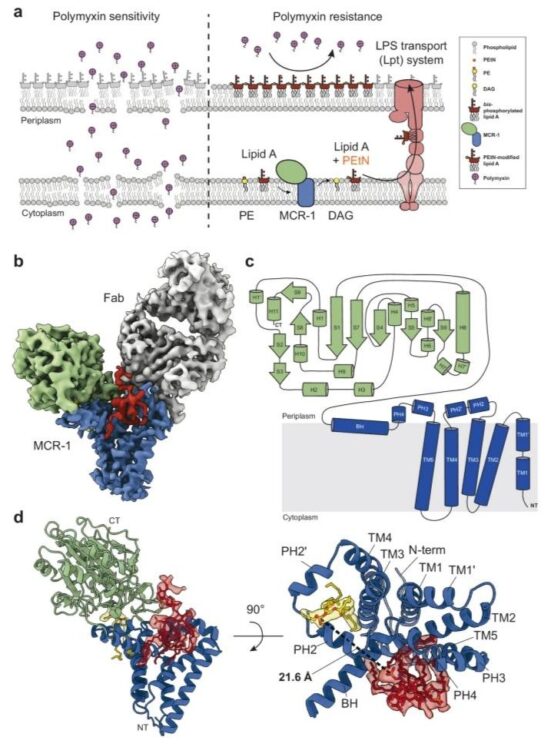
Mechanistic basis of antimicrobial resistance mediated by the phosphoethanolamine transferase MCR-1 Journal Article
In: Nat Commun, vol. 16, no. 1, pp. 10516, 2025, ISSN: 2041-1723.
@article{pmid41298376,
title = {Mechanistic basis of antimicrobial resistance mediated by the phosphoethanolamine transferase MCR-1},
author = {Allen P Zinkle and Mariana Bunoro Batista and Carmen M Herrera and Satchal K Erramilli and Brian Kloss and Khuram U Ashraf and Kamil Nosol and Guozhi Zhang and Rosemary J Cater and Michael T Marty and Anthony A Kossiakoff and M Stephen Trent and Rie Nygaard and Phillip J Stansfeld and Filippo Mancia},
doi = {10.1038/s41467-025-65515-3},
issn = {2041-1723},
year = {2025},
date = {2025-11-01},
urldate = {2025-11-01},
journal = {Nat Commun},
volume = {16},
number = {1},
pages = {10516},
abstract = {Polymyxins are used to treat infections caused by multidrug-resistant Gram-negative bacteria. They are cationic peptides that target the negatively charged lipid A component of lipopolysaccharides, disrupting the outer membrane and lysing the cell. Polymyxin resistance is conferred by inner-membrane enzymes, such as phosphoethanolamine transferases, which add positively charged phosphoethanolamine to lipid A. Here, we present the structure of MCR-1, a plasmid-encoded phosphoethanolamine transferase, in its liganded form. The phosphatidylethanolamine donor substrate is bound near the active site in the periplasmic domain, and lipid A is bound over 20 Å away, within the transmembrane region. Integrating structural, biochemical, and drug-resistance data with computational analyses, we propose a two-state model in which the periplasmic domain rotates to bring the active site to lipid A, near the preferential phosphate modification site for MCR-1. This enzymatic mechanism may be generally applicable to other phosphoform transferases with large, globular soluble domains.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Romane, Ksenija; Peteani, Giulia; Mukherjee, Somnath; Kowal, Julia; Rossi, Lorenzo; Hou, Jingkai; Kossiakoff, Anthony A; Lemmin, Thomas; Locher, Kaspar P
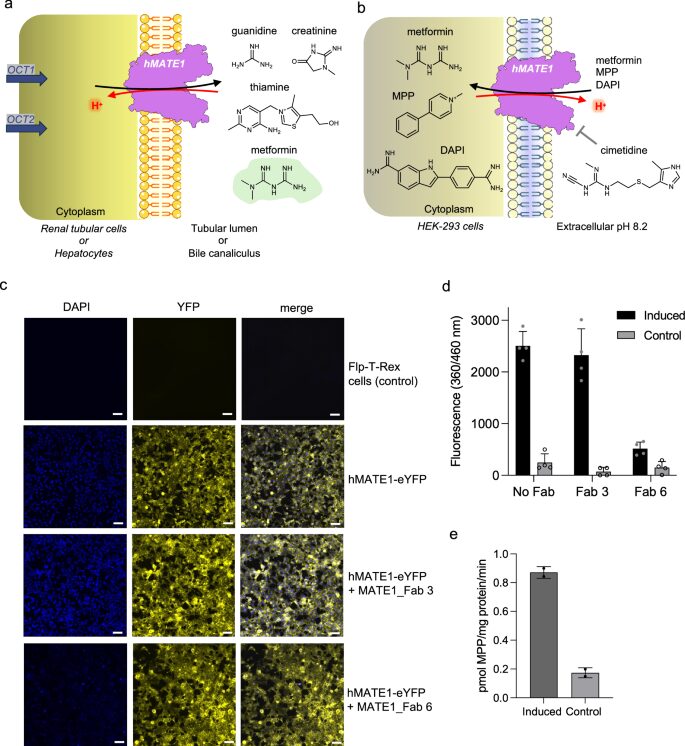
Structural basis of drug recognition by human MATE1 transporter Journal Article
In: Nat Commun, vol. 16, no. 1, pp. 9444, 2025, ISSN: 2041-1723.
@article{pmid41145429,
title = {Structural basis of drug recognition by human MATE1 transporter},
author = {Ksenija Romane and Giulia Peteani and Somnath Mukherjee and Julia Kowal and Lorenzo Rossi and Jingkai Hou and Anthony A Kossiakoff and Thomas Lemmin and Kaspar P Locher},
doi = {10.1038/s41467-025-64490-z},
issn = {2041-1723},
year = {2025},
date = {2025-10-28},
urldate = {2025-10-01},
journal = {Nat Commun},
volume = {16},
number = {1},
pages = {9444},
abstract = {Human MATE1 (multidrug and toxin extrusion protein 1) is highly expressed in the kidney and liver, where it mediates the final step in the excretion of a broad range of cationic drugs, including the antidiabetic drug metformin, into the urine and bile. This transport process is essential for drug clearance and also affects therapeutic efficacy. To understand the molecular basis of drug recognition by hMATE1, we determined cryo-electron microscopy structures of the transporter in complex with the substrates 1-methyl-4-phenylpyridinium (MPP) and metformin and with the inhibitor cimetidine. The structures reveal a shared binding site located in a negatively charged pocket in the C-lobe of the protein. We functionally validated key interactions using radioactivity-based cellular uptake assays using hMATE1 mutants. Molecular dynamics simulations provide insights into the different binding modes and dynamic behaviour of the ligands within the pocket. Collectively, these findings define the structural basis of hMATE1 substrate specificity and shed light on its role in drug transport and drug-drug interactions.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Zhao, Fangzhu; Wu, Yan; Schaefer, Kaitlin; Zhang, Yun; Miao, Kun; Yao, Zi; Ganjave, Snehal D; Kumru, Kaan; Peters-Clarke, Trenton M; Inague, Alex; Olzmann, James A; Leung, Kevin K; Wells, James A
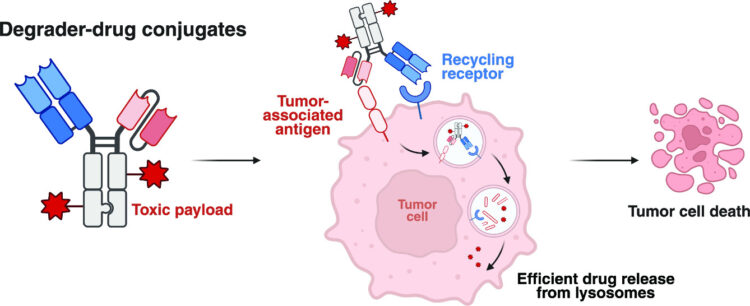
Hijacking Extracellular Targeted Protein Degrader-Drug Conjugates for Enhanced Drug Delivery Journal Article
In: J Am Chem Soc, 2025, ISSN: 1520-5126.
@article{pmid41091621,
title = {Hijacking Extracellular Targeted Protein Degrader-Drug Conjugates for Enhanced Drug Delivery},
author = {Fangzhu Zhao and Yan Wu and Kaitlin Schaefer and Yun Zhang and Kun Miao and Zi Yao and Snehal D Ganjave and Kaan Kumru and Trenton M Peters-Clarke and Alex Inague and James A Olzmann and Kevin K Leung and James A Wells},
doi = {10.1021/jacs.5c15047},
issn = {1520-5126},
year = {2025},
date = {2025-10-15},
urldate = {2025-10-01},
journal = {J Am Chem Soc},
abstract = {Antibody-based therapeutics encompass diverse modalities for targeting tumor cells. Among these, antibody-drug conjugates (ADCs) and extracellular targeted protein degradation (eTPD) specifically depend on efficient lysosomal trafficking for activity. A major limitation of ADCs is their reliance on antigens with efficient internalization, while eTPD approaches, although capable of trafficking diverse targets to lysosomes, lack cytotoxic potency. To address this, we developed degrader-drug conjugates (DDCs), leveraging the endocytic and recycling activities of eTPD to enhance lysosomal delivery. We utilized fast internalizers, the low-density lipoprotein receptor (LDLR) and the chemokine receptor (CXCR7), to enhance lysosomal delivery. LDLR-based degraders enabled efficient and selective degradation of diverse extracellular membrane proteins, while DDCs with cytotoxic payload enhanced cytotoxicity compared to conventional ADCs . This dual modality addresses key challenges of inadequate internalization in conventional ADCs and cytotoxic potency in current eTPD strategies. Our findings demonstrate that DDCs provide additional optionality for developing next-generation antibody therapeutics with broader utility and improved efficacy in cancer treatment.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Mohona, Surovi; Shakya, Anil K; Singh, Suruchi; Kearns, Fiona L; Jemison, Kezia; Erramilli, Satchal; Dey, Debajit; Qing, Enya; Jennings, Benjamin C; Doray, Balraj; Kossiakoff, Anthony A; Amaro, Rommie E; Klose, Thomas; Gallagher, Tom; Hasan, S Saif
An unconventional HxD motif orchestrates coatomer-dependent coronavirus morphogenesis Journal Article
In: bioRxiv, 2025, ISSN: 2692-8205.
@article{pmid41279991,
title = {An unconventional HxD motif orchestrates coatomer-dependent coronavirus morphogenesis},
author = {Surovi Mohona and Anil K Shakya and Suruchi Singh and Fiona L Kearns and Kezia Jemison and Satchal Erramilli and Debajit Dey and Enya Qing and Benjamin C Jennings and Balraj Doray and Anthony A Kossiakoff and Rommie E Amaro and Thomas Klose and Tom Gallagher and S Saif Hasan},
doi = {10.1101/2025.10.16.682669},
issn = {2692-8205},
year = {2025},
date = {2025-10-01},
urldate = {2025-10-01},
journal = {bioRxiv},
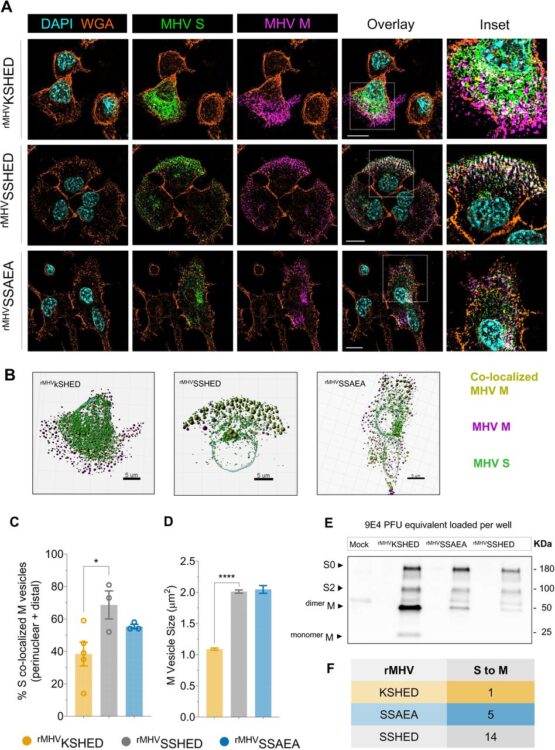
abstract = {Assembly of infectious coronaviruses requires spike (S) protein trafficking by host coatomer, typically via a dibasic signal in the S cytoplasmic tail. However, the human embecoviruses HKU1 and OC43, as well as the model virus MHV, lack this motif. Here we identify a conserved His-x-Asp (HxD) sequence that functions as an unconventional coatomer-binding signal. Structural and biochemical analyses show that the MHV HxD motif engages coatomer subunits through distinct conformations, while cellular imaging demonstrates its role in directing S to assembly sites with the viral M-protein. Disruption of HxD-coatomer interactions impairs S incorporation and provokes compensatory viral adaptations, including emergence of a canonical dibasic motif or mutations in M-protein. Electron microscopy further reveals profound alterations in virion surface architecture. These findings uncover HxD as a previously unrecognized coatomer-targeting motif, highlighting an unexpected flexibility in coronavirus assembly pathways and broadening understanding of the cellular machinery that shapes coronavirus morphogenesis.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Adams, Jarrett J.; Blazer, Levi L.; Chung, Jacky; Karimi, Minoo; Davidson, Taylor; Bruce, Heather A.; Singer, Alexander U.; Yang, Ning; Cardarelli, Lia; Pot, Isabelle; Colombo, Luigi; Huang, Lily Jun‐shen; Ma, Yue; Michnick, Stephen W.; Moe, Orson W.; Sidhu, Sachdev S.
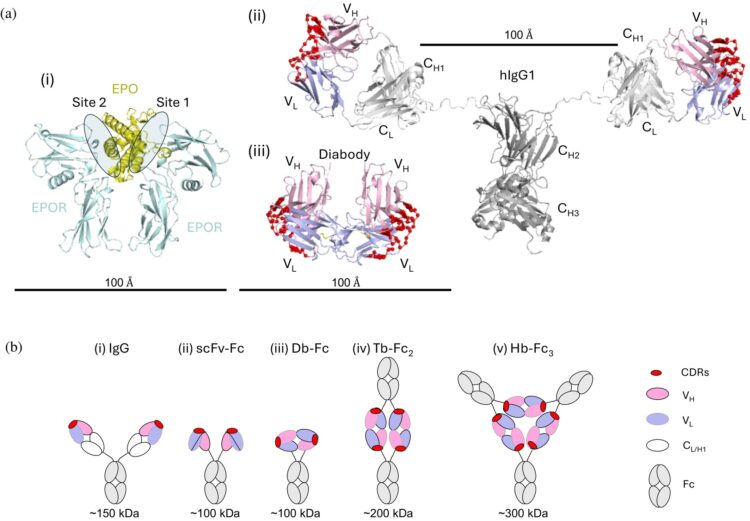
An asymmetric tetrabody is a potent and efficacious agonist of the erythropoietin receptor in vitro and in vivo Journal Article
In: Protein Science, vol. 34, no. 10, 2025, ISSN: 1469-896X.
@article{Adams2025,
title = {An asymmetric tetrabody is a potent and efficacious agonist of the erythropoietin receptor in vitro and in vivo},
author = {Jarrett J. Adams and Levi L. Blazer and Jacky Chung and Minoo Karimi and Taylor Davidson and Heather A. Bruce and Alexander U. Singer and Ning Yang and Lia Cardarelli and Isabelle Pot and Luigi Colombo and Lily Jun‐shen Huang and Yue Ma and Stephen W. Michnick and Orson W. Moe and Sachdev S. Sidhu},
doi = {10.1002/pro.70292},
issn = {1469-896X},
year = {2025},
date = {2025-09-17},
urldate = {2025-10-00},
journal = {Protein Science},
volume = {34},
number = {10},
publisher = {Wiley},
abstract = {Abstract: Erythropoietin (EPO) initiates EPO receptor (EPOR) signaling in hematopoietic cells by binding to an asymmetric EPOR dimer through two different sites. We engineered dimeric diabody‐Fc (Db‐Fc) fusion proteins that appeared to act as potent agonists of human EPOR in cell proliferation assays. However, detailed analysis of their oligomeric forms revealed that the predominant Db‐Fc species bound EPOR with high affinity but failed to induce cell proliferation. Instead, a minor oligomeric form, identified as a putative tetrabody (Tb) fused to two Fc domains (Tb‐Fc), proved to be the minimal active form. The existence of a tetrameric agonist was further supported by crystallography, which revealed an asymmetric Tb structure. Additionally, the structure of an antigen‐binding fragment (Fab) bound to EPOR revealed an epitope distinct from the EPO binding sites, and structural modeling showed that engagement of two of the four binding sites on the Tb could form an asymmetric EPOR dimer nearly identical to the active conformation recruited by EPO. In a knock‐in mouse model, where mouse EPOR was replaced by human EPOR, purified Tb‐Fc stimulated erythropoiesis with greater potency, efficacy, and duration than darbepoetin, a recombinant EPO that is the leading therapeutic erythropoiesis‐stimulating agent (ESA). Collectively, these findings demonstrate that asymmetric tetravalent antibodies such as Tb‐Fc represent promising next‐generation ESAs that provide enhanced potency, efficacy, and durability. Moreover, they may reduce the oncogenic and cardiovascular risks associated with the pleiotropy of EPO.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Wadhwa, Anju; Johnson, Haley; Bobba, Kondapa Naidu; Bidkar, Anil P; Mayne, Ellis; Rampersaud, Sham; Mandal, Kamal; Barpanda, Abhilash; Prudhvi, Sanjana; Kang, Amrik S; Greenland, Nancy; Balitzer, Dana; Peter, Robin; Raveendran, Athira; Naik, Shubhankar; Basak, Megha; Kasap, Corynn; Huebner, Juwita; Alvarez, Marina Lopez; Lee, Sanghee; Steri, Veronica; Adams, Jarret J; Sidhu, Sachdev S; Wilson, David M; Seo, Youngho; VanBrocklin, Henry F; Logan, Aaron C; Wiita, Arun P; Flavell, Robert R
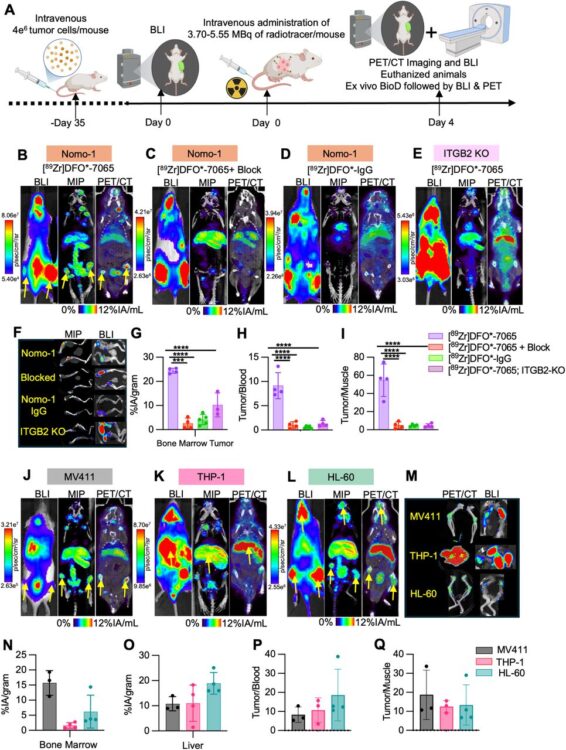
Effective imaging and treatment of Acute Myeloid Leukemia with radiotheranostics targeting the activated conformation of integrin-βeta2 Journal Article
In: bioRxiv, 2025, ISSN: 2692-8205.
@article{pmid40909732,
title = {Effective imaging and treatment of Acute Myeloid Leukemia with radiotheranostics targeting the activated conformation of integrin-βeta2},
author = {Anju Wadhwa and Haley Johnson and Kondapa Naidu Bobba and Anil P Bidkar and Ellis Mayne and Sham Rampersaud and Kamal Mandal and Abhilash Barpanda and Sanjana Prudhvi and Amrik S Kang and Nancy Greenland and Dana Balitzer and Robin Peter and Athira Raveendran and Shubhankar Naik and Megha Basak and Corynn Kasap and Juwita Huebner and Marina Lopez Alvarez and Sanghee Lee and Veronica Steri and Jarret J Adams and Sachdev S Sidhu and David M Wilson and Youngho Seo and Henry F VanBrocklin and Aaron C Logan and Arun P Wiita and Robert R Flavell},
doi = {10.1101/2025.08.21.671206},
issn = {2692-8205},
year = {2025},
date = {2025-08-01},
urldate = {2025-08-01},
journal = {bioRxiv},
abstract = {There remains an unmet clinical need for improved treatment strategies in Acute Myeloid Leukemia (AML). Although radiopharmaceutical therapies targeting non-cancer-selective antigens have shown promise in AML, their clinical utility is often limited by prolonged bone marrow suppression. Using a unique proteomics-based strategy, we recently identified the active conformation of integrin-β2 (aITGB2) as a novel, tumor-selective target for AML. Importantly, this conformational epitope is expressed widely on AML cells but minimally on normal marrow progenitors/healthy tissues. Here we first confirmed widespread aITGB2 expression on AML tumors that was largely independent of tumor genotype or prior therapeutic regimen. We developed diagnostic and therapeutic radiopharmaceuticals targeting aITGB2 utilizing a conformation-specific antibody (clone 7065). PET/CT imaging with Zr and Ce-labeled 7065 in AML models revealed high target-mediated uptake, greater than that compared to standard of care [ F]-FDG. PET/CT imaging with [ Zr]DFO*-7065 showed reduced binding to normal bone marrow and immune cells in humanized immune system mice compared to [ Zr]DFO*-anti-CD33. For therapy, we developed [ Ac]Macropa-PEG -7065 using an optimized chelator-linker combination. Treatment with [ Ac]Macropa-PEG -7065 in Nomo-1 and PDX AML disseminated models delayed tumor growth and improved overall survival compared to controls, including [ Ac]DOTA-anti-CD33, a clinical stage-radioimmunotherapy under evaluation in AML. Relapsed tumors demonstrated persistent aITGB2 expression, supporting continued development of fractionated dosing schemes, and proteomics analysis indicated activation of TCA cycle and carbon metabolism pathways, consistent with therapy-induced stress responses. These findings highlight [ Zr]DFO*-7065 and [ Ac]Macropa-7065 as a promising aITGB2-targeted theranostic pair with potential for imaging and treatment in future clinical translation.nnONE SENTENCE SUMMARY: This study demonstrates promising preclinical efficacy of aITGB2-targeted radiotheranostics for selective imaging and therapy in AML.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Cottom, Claire Overly; Heinz, Eva; Erramilli, Satchal; Kossiakoff, Anthony; Slade, Daniel J; Noinaj, Nicholas
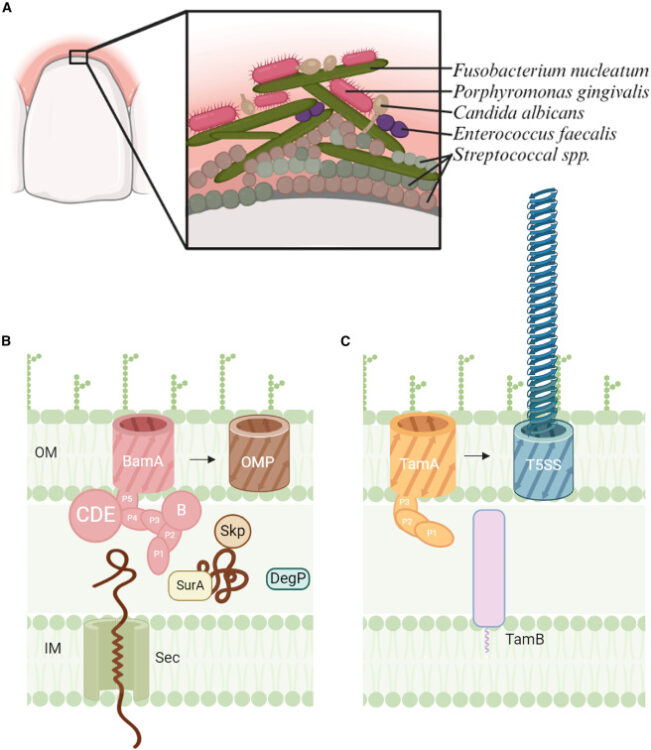
Characterization of the OMP biogenesis machinery in Fusobacterium nucleatum Journal Article
In: Structure, 2025, ISSN: 1878-4186.
@article{pmid40897170,
title = {Characterization of the OMP biogenesis machinery in Fusobacterium nucleatum},
author = {Claire Overly Cottom and Eva Heinz and Satchal Erramilli and Anthony Kossiakoff and Daniel J Slade and Nicholas Noinaj},
doi = {10.1016/j.str.2025.08.008},
issn = {1878-4186},
year = {2025},
date = {2025-08-01},
urldate = {2025-08-01},
journal = {Structure},
abstract = {F. nucleatum is a Gram-negative bacteria that causes oral infections and is linked to colorectal cancer. Pathogenicity relies on a type of β-barrel outer membrane protein (OMP) called an autotransporter. The biogenesis of OMPs is typically mediated by the barrel assembly machinery (BAM) complex. In this study, we investigate the evolution, composition, and structure of the OMP biogenesis machinery in F. nucleatum. Our bioinformatics and proteomics analyses indicate that OMP biogenesis in F. nucleatum is mediated solely by the core component BamA. The structure of FnBamA highlights distinct features, including four POTRA domains and a C-terminal 16-stranded β-barrel domain observed as an inverted dimer. FnBamA represents the original composition of the assembly machinery, and a duplication event that resulted in BamA and TamA occurred after the split of other lineages, including the Proteobacteria, from the Fusobacteria. FnBamA, therefore, likely serves a singular role in the biogenesis of all OMPs.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
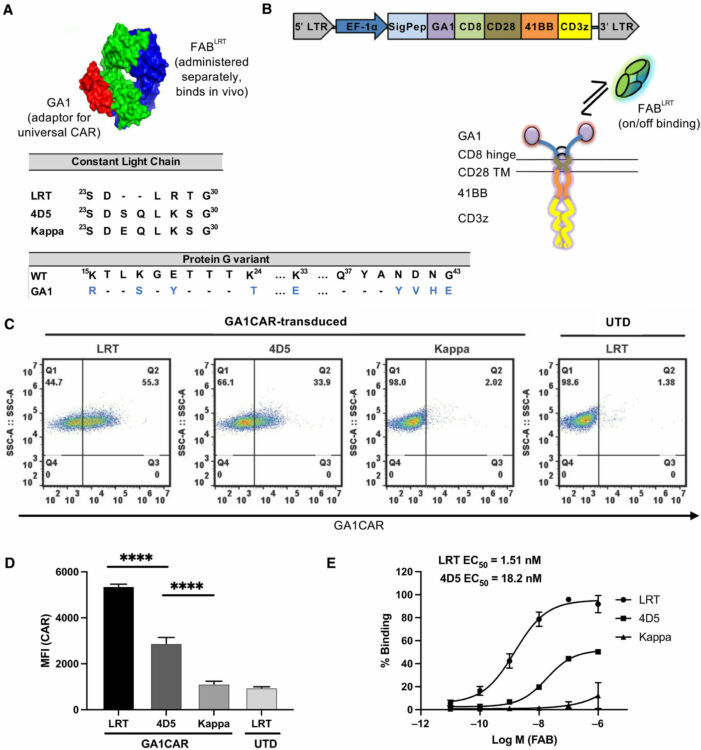
Arina, Ainhoa; Arauz, Edwin; Masoumi, Elham; Warzecha, Karolina W; Sääf, Annika; Widło, Łukasz; Slezak, Tomasz; Zieminska, Aleksandra; Dudek, Karolina; Schaefer, Zachary P; Lecka, Maria; Usatyuk, Svitlana; Weichselbaum, Ralph R; Kossiakoff, Anthony A
A universal chimeric antigen receptor (CAR)-fragment antibody binder (FAB) split system for cancer immunotherapy Journal Article
In: Sci Adv, vol. 11, no. 27, pp. eadv4937, 2025, ISSN: 2375-2548.
@article{pmid40614208,
title = {A universal chimeric antigen receptor (CAR)-fragment antibody binder (FAB) split system for cancer immunotherapy},
author = {Ainhoa Arina and Edwin Arauz and Elham Masoumi and Karolina W Warzecha and Annika Sääf and Łukasz Widło and Tomasz Slezak and Aleksandra Zieminska and Karolina Dudek and Zachary P Schaefer and Maria Lecka and Svitlana Usatyuk and Ralph R Weichselbaum and Anthony A Kossiakoff},
doi = {10.1126/sciadv.adv4937},
issn = {2375-2548},
year = {2025},
date = {2025-07-01},
urldate = {2025-07-01},
journal = {Sci Adv},
volume = {11},
number = {27},
pages = {eadv4937},
abstract = {Chimeric antigen receptor (CAR) T cell therapy has shown extraordinary results in treating hematological cancer but faces challenges like antigen loss, toxicity, and complex manufacturing. Universal and modular CAR constructs offer improved flexibility, safety, and cost-effectiveness over conventional CAR constructs. We present a CAR-fragment antibody binder (Fab) platform on the basis of an engineered protein G variant (GA1) and Fab scaffolds. Expression of GA1CAR on human CD8 T cells leads to antigen recognition and T cell effector function that can be modulated according to the affinity of the CAR for the Fab and of the Fab for the target. GA1CAR T cells can recognize multiple Fab-antigen pairs on breast and ovarian cancer cell lines. Adoptively transferred GA1CAR T cells control tumors in breast cancer xenograft models, and their targeting can be quickly redirected using different Fabs. This versatile "plug-and-play" CAR T platform has potential for application in personalized therapy, preventing antigen loss variant escape, decreasing toxicity, and increasing access.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}