Publications: 2026
2026
Alexander, J Andrew N; Chen, Shu-Yu; Mukherjee, Somnath; de Capitani, Mario; Irobalieva, Rossitza N; Rossi, Lorenzo; Agrawal, Parth; Kowal, Julia; Meirelles, Matheus A; Aebi, Markus; Reymond, Jean-Louis; Kossiakoff, Anthony A; Riniker, Sereina; Locher, Kaspar P
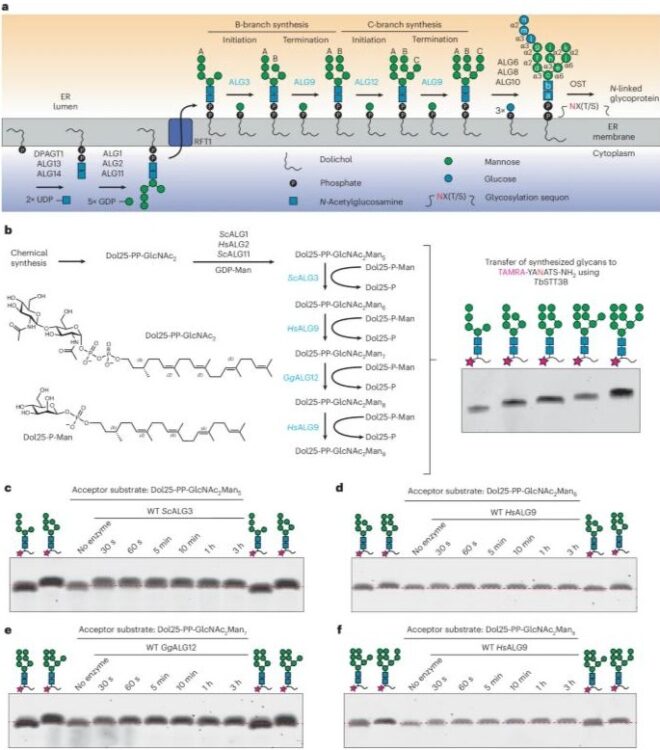
Structures of ALG3/9/12 reveal the assembly logic of the N-glycan oligomannose core Journal Article
In: Nat Chem Biol, 2026, ISSN: 1552-4469.
@article{pmid41807832,
title = {Structures of ALG3/9/12 reveal the assembly logic of the N-glycan oligomannose core},
author = {J Andrew N Alexander and Shu-Yu Chen and Somnath Mukherjee and Mario de Capitani and Rossitza N Irobalieva and Lorenzo Rossi and Parth Agrawal and Julia Kowal and Matheus A Meirelles and Markus Aebi and Jean-Louis Reymond and Anthony A Kossiakoff and Sereina Riniker and Kaspar P Locher},
doi = {10.1038/s41589-026-02164-7},
issn = {1552-4469},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {Nat Chem Biol},
abstract = {Asparagine-linked glycans are essential for the maturation and function of most eukaryotic secretory proteins. The biosynthesis and transfer of dolichylpyrophosphate-anchored GlcNAcManGlc glycan is a highly conserved process occurring in the endoplasmic reticulum (ER) membrane and involving over a dozen membrane proteins whose dysfunction is linked to congenital disorders of glycosylation (CDGs). Three membrane-integral mannosyltransferases, ALG3, ALG9 and ALG12, mediate four consecutive mannosylation reactions that convert GlcNAcMan to GlcNAcMan. Here, using chemoenzymatically synthesized lipid-linked glycan donor and acceptor analogs, we recapitulated this biosynthetic pathway in vitro. High-resolution cryo-electron microscopy structures of pseudo-Michaelis complexes of each step revealed how the branched glycan is accurately synthesized and unwanted side products are averted. Molecular dynamics simulations and mutagenesis studies uncovered a subtle but precise mechanism selecting the dolichylphosphomannose donor substrate over dolichylphosphoglucose, which is also present in the ER membrane. Our results also provide mechanistic explanations for enzyme dysfunction in CDGs and offer opportunities for N-glycan engineering.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Delaveris, Corleone S; Loudermilk, Rita P; Pandey, Apurva; Remesh, Soumya G; Peters-Clarke, Trenton M; Ganjave, Snehal D; Dougherty, William J N; Delavan, Henry M; Wang, Chunyue; Ling, Jesse; Serrano, Juan Antonio Camara; Salangsang, Fernando; Ding, Chien-Kuang Cornelia; Greenland, Nancy; Chu, Carissa E; Porten, Sima; Steri, Veronica; Chou, Jonathan; Evans, Michael J; Leung, Kevin K; Wells, James A
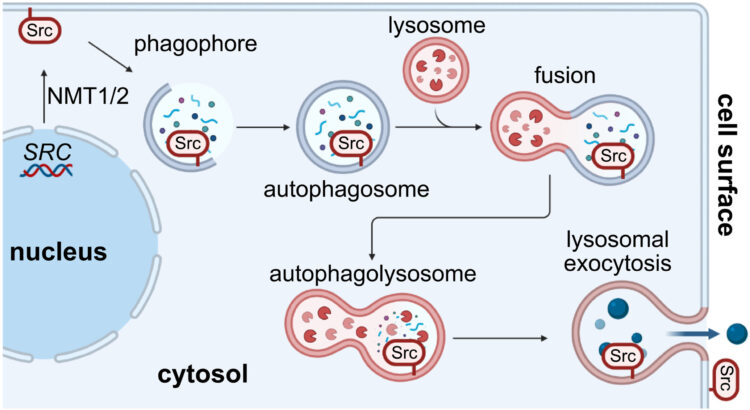
Autophagolysosomal exocytosis inverts Src kinase onto the cell surface in cancer Journal Article
In: Science, vol. 391, no. 6790, pp. eaec1778, 2026, ISSN: 1095-9203.
@article{pmid41818370,
title = {Autophagolysosomal exocytosis inverts Src kinase onto the cell surface in cancer},
author = {Corleone S Delaveris and Rita P Loudermilk and Apurva Pandey and Soumya G Remesh and Trenton M Peters-Clarke and Snehal D Ganjave and William J N Dougherty and Henry M Delavan and Chunyue Wang and Jesse Ling and Juan Antonio Camara Serrano and Fernando Salangsang and Chien-Kuang Cornelia Ding and Nancy Greenland and Carissa E Chu and Sima Porten and Veronica Steri and Jonathan Chou and Michael J Evans and Kevin K Leung and James A Wells},
doi = {10.1126/science.aec1778},
issn = {1095-9203},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {Science},
volume = {391},
number = {6790},
pages = {eaec1778},
abstract = {Overexpression of the proto-oncogene Src is common to a wide variety of cancers. In this work, we found that Src is noncanonically translocated and inverted onto the cell surface in cancer, both in vitro and in vivo. We identified autophagolysosomal exocytosis (ALE) as a secretory mechanism prominent in cancer cell lines. Src represents the prototypical example of a family of membrane-anchored proteins that are transported by this process. Furthermore, this extracellular membrane-associated Src (eSrc) was found in primary tumors, and anti-Src antibody-based therapies mediated tumor cell killing in cell culture systems and in mouse xenograft models. Thus, intracellular -myristoylated proteins, prototypically Src, can be topologically inverted onto the cell surface in cancer and targeted with antibody therapeutics.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Slezak, Tomasz; O'Leary, Kelly M; Avella, Tanya Guevara; Musial, Natalia; Li, Jinyang; Andrzejczak, Anna; Scott, Elizabeth F; Le, Duc Anh; Kossiakoff, Anthony A
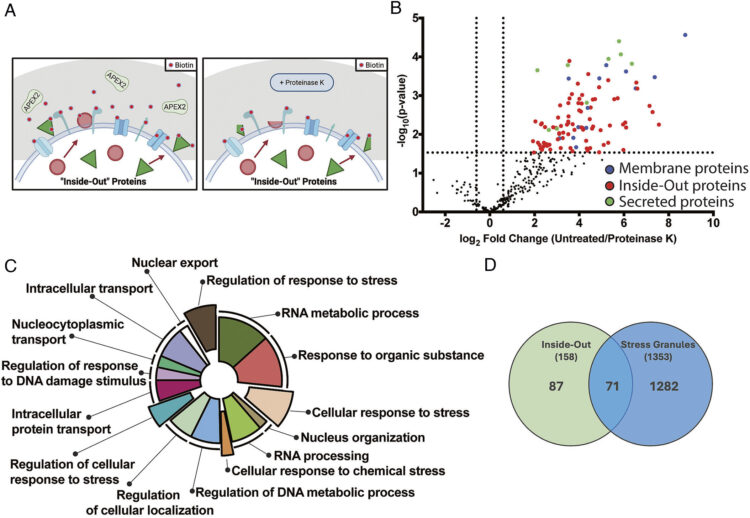
Dynamic translocation of Inside-Out proteins to the cell surface underlies cellular adaptation to cancer-induced stress Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 13, pp. e2529493123, 2026, ISSN: 1091-6490.
@article{pmid41875162,
title = {Dynamic translocation of Inside-Out proteins to the cell surface underlies cellular adaptation to cancer-induced stress},
author = {Tomasz Slezak and Kelly M O'Leary and Tanya Guevara Avella and Natalia Musial and Jinyang Li and Anna Andrzejczak and Elizabeth F Scott and Duc Anh Le and Anthony A Kossiakoff},
doi = {10.1073/pnas.2529493123},
issn = {1091-6490},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {13},
pages = {e2529493123},
abstract = {Inside-Out (I-O) protein display, the noncanonical surface localization of intracellular proteins, represents an underexplored feature of tumor cell biology. Here, we map the molecular landscape and trafficking mechanisms that control the presentation of I-O proteins on cancer cell membranes. Employing APEX2-mediated proximity biotinylation and a custom antibody generation and validation platform, we identified approximately 140 high-confidence I-O proteins, primarily ribosomal, proteasomal, chaperone, and translation factors, notably enriched in protein families associated with stress-response pathways. Validation of 500 antibodies encompassing 40 I-O targets across seven tumor cell lines confirmed selective and robust surface localization, while in vivo imaging in mouse xenografts demonstrated pronounced and tumor-specific antibody accumulation. I-O proteins were absent on peripheral blood mononuclear cells (PBMCs) and in normal tissues, indicating cancer cell selectivity. Functional analyses revealed that I-O protein tethering to the membrane is dependent on heparan sulfate interactions; enzymatic removal of these glycans led to the clearance of I-O proteins from the cell surface. Notably, the removed proteins returned to baseline levels within 6 h, indicating a dynamic balance related to Endoplasmic Reticulum (ER)-Golgi trafficking and cellular stress. Nearly half of these I-O proteins overlapped with known stress granule (SG) components; however, stress elements that promote SG formation do not similarly affect surface display of I-O proteins. Furthermore, I-O proteins are present on standard cancer cell lines under lower stress levels needed to induce SG formation, suggesting parallel yet mechanistically distinct aspects of the stress response. These findings position I-O display as a paradigm in protein trafficking, different from traditional secretion pathways and closely linked to stress response.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Anderson, Kyle J; Lee, Melody S; Sevillano, Natalia; Chen, Gang; Hornsby, Michael J; Sidhu, Sachdev S; Craik, Charles S
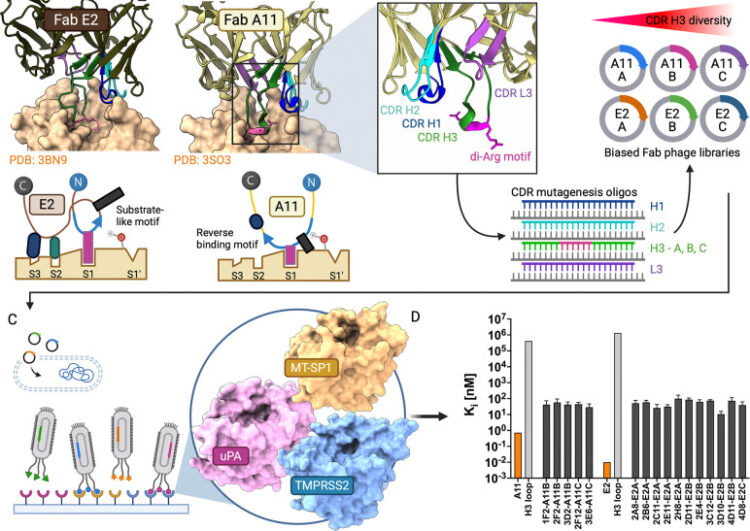
Structural Basis of Serine Protease Inhibition by Antibodies from Biased Fab Phage-Display Libraries Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid41959119,
title = {Structural Basis of Serine Protease Inhibition by Antibodies from Biased Fab Phage-Display Libraries},
author = {Kyle J Anderson and Melody S Lee and Natalia Sevillano and Gang Chen and Michael J Hornsby and Sachdev S Sidhu and Charles S Craik},
doi = {10.64898/2026.03.12.711446},
issn = {2692-8205},
year = {2026},
date = {2026-03-01},
urldate = {2026-03-01},
journal = {bioRxiv},
abstract = {Biased Fab phage-display libraries were designed to determine whether inhibitory CDR H3 motifs from potent anti-matriptase antibodies could be transferred to target homologous serine proteases. Using reverse-binding and substrate-like H3 motifs from parental clones A11 and E2 as templates, six synthetic libraries with 10 diversity were constructed. Selection against matriptase identified sixteen inhibitors with sub-100 nM potency, representing 100,000-fold improvement over circularized H3 loops alone. Selection against TMPRSS2, a serine protease implicated in viral entry and prostate cancer with 43% sequence identity to matriptase, yielded binders with micromolar inhibitory potency. Selection against urokinase plasminogen activator (uPA, 35% identity) identified binders that adopted a substrate-like CDR H3 binding mode in our structural models. Across all reference structures, including the separately identified uPA inhibitor AB2 (PDB: 9PYF, deposited with this work), benchmarking of five co-folding methods and rigid-body docking showed that co-folding consistently achieved acceptable to high quality DockQ scores, outperforming traditional docking and capturing the recognition of key active site determinants. Ensemble predictions of mutational binding energy changes (ΔΔ) using these models identified key paratope-epitope interactions, with predictions validated through mutagenesis. This work establishes a framework integrating biased antibody libraries with computational structure prediction and analysis for targeting conserved protease epitopes.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Lin, Zhi; Ngo, Wayne; Chou, Yu-Ting; Wu, Harry; Susa, Katherine J; Jun, Young-Wook; Bivona, Trever G; Doudna, Jennifer A; Wells, James A
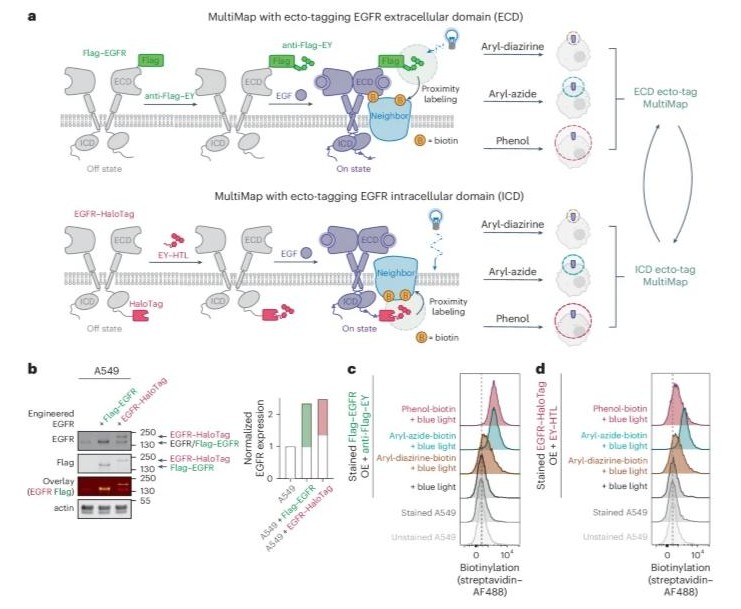
Temporal photoproximity labeling of ligand-activated EGFR neighborhoods using MultiMap Journal Article
In: Nat Chem Biol, vol. 22, no. 2, pp. 192–204, 2026, ISSN: 1552-4469.
@article{pmid41254216,
title = {Temporal photoproximity labeling of ligand-activated EGFR neighborhoods using MultiMap},
author = {Zhi Lin and Wayne Ngo and Yu-Ting Chou and Harry Wu and Katherine J Susa and Young-Wook Jun and Trever G Bivona and Jennifer A Doudna and James A Wells},
doi = {10.1038/s41589-025-02076-y},
issn = {1552-4469},
year = {2026},
date = {2026-02-01},
urldate = {2025-11-01},
journal = {Nat Chem Biol},
volume = {22},
number = {2},
pages = {192--204},
abstract = {Photoproximity labeling proteomics (PLP) methods have recently shown that cell surface receptors can form lateral interactome networks. Here, we present a paired set of PLP workflows that dynamically track neighborhood changes for oncogenic epidermal growth factor receptor (EGFR) over time, both outside and inside of cells. We achieved this by augmenting the multiscale PLP workflow we call MultiMap, where three photoprobes with different labeling ranges were photoactivated by one photocatalyst, eosin Y, anchored extracellularly and intracellularly on EGFR. We identified hundreds of neighboring proteins that changed within minutes to over 1 h after the addition of EGF. These neighborhoods reveal dynamic interactomes during early, middle and late signaling that drive phosphorylation, internalization, degradation and transcriptional regulation. This rapid 'molecular photographic' labeling approach provides snapshots of signaling neighborhoods, revealing their dynamic nature and potential for drug targeting.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Adams, Jarrett J; Blazer, Levi L; Chung, Jacky; Karimi, Minoo; Davidson, Taylor; Blair, Bailey; Waddle, Carlos; Hokanson, Craig A; Bruce, Heather A; Singer, Alexander U; Tombak, Eva-Maria; Gildemann, Kiira; Tamberg, Nele; Kiiver, Kaja; Ustav, Mart; Ma, Yue; Colombo, Luigi; Huang, Lily Jun-Shen; Michnick, Stephen W; Moe, Orson W; Sidhu, Sachdev S
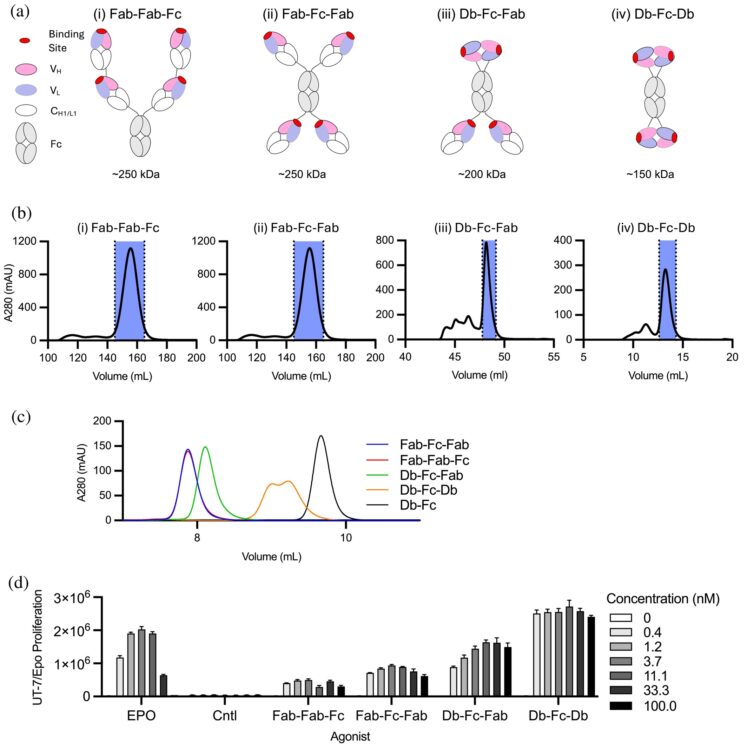
Tetravalent antibodies are more potent and efficacious erythropoiesis-stimulating agents than erythropoietin in vivo Journal Article
In: Protein Sci, vol. 35, no. 2, pp. e70462, 2026, ISSN: 1469-896X.
@article{pmid41556618,
title = {Tetravalent antibodies are more potent and efficacious erythropoiesis-stimulating agents than erythropoietin in vivo},
author = {Jarrett J Adams and Levi L Blazer and Jacky Chung and Minoo Karimi and Taylor Davidson and Bailey Blair and Carlos Waddle and Craig A Hokanson and Heather A Bruce and Alexander U Singer and Eva-Maria Tombak and Kiira Gildemann and Nele Tamberg and Kaja Kiiver and Mart Ustav and Yue Ma and Luigi Colombo and Lily Jun-Shen Huang and Stephen W Michnick and Orson W Moe and Sachdev S Sidhu},
doi = {10.1002/pro.70462},
issn = {1469-896X},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {Protein Sci},
volume = {35},
number = {2},
pages = {e70462},
abstract = {Recent studies have shown that tetravalent antibodies are potent and efficacious agonists of the erythropoietin (EPO) receptor (EPOR) both in vitro and in vivo. To identify antibody-based erythropoiesis-stimulating agents (ESAs) with therapeutic potential, we evaluated various tetravalent antibody formats for EPOR agonism and key biophysical properties necessary for biologic drug development. We identified two distinct tetravalent antibody formats that strongly stimulated the growth of UT7/Epo cells, which rely on EPOR signaling for proliferation. Moreover, one of these formats exhibited ideal biophysical characteristics for drug development. This format consisted of a diabody (Db) and two antigen-binding fragment (Fab) arms fused to the N- and C-termini of an Fc domain, respectively, to form a tetravalent Db-Fc-Fab (EPRA-0322). In a mouse model expressing the human EPOR, EPRA-0322 induced erythropoiesis with greater potency, efficacy, and duration than darbepoetin, a hyperglycosylated EPO currently used in clinical practice. These findings highlight tetravalent antibodies, and the Db-Fc-Fab format in particular, as promising next-generation ESAs suitable for large-scale production and clinical use.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Erramilli, Satchal K; Nosol, Kamil; Pietrzak-Lichwa, Krzysztof; Schmandt, Nicolaus; Li, Tian; Tokarz, Piotr; Hou, Jingkai; Zhao, Minglei; Perozo, Eduardo; Kossiakoff, Anthony A
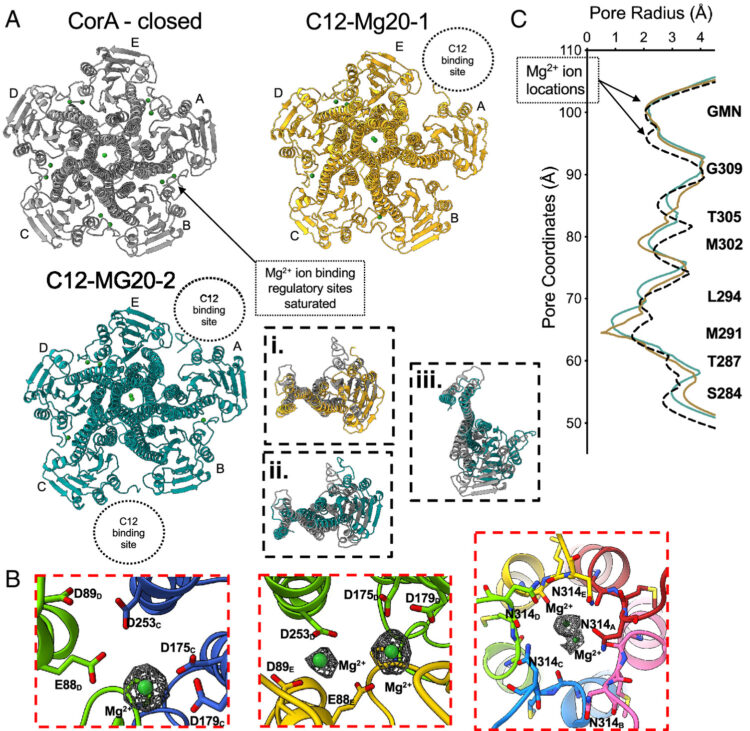
Conformational ensembles of the magnesium channel CorA reveal structural basis for channel gating Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 8, pp. e2512532123, 2026, ISSN: 1091-6490.
@article{pmid41701836,
title = {Conformational ensembles of the magnesium channel CorA reveal structural basis for channel gating},
author = {Satchal K Erramilli and Kamil Nosol and Krzysztof Pietrzak-Lichwa and Nicolaus Schmandt and Tian Li and Piotr Tokarz and Jingkai Hou and Minglei Zhao and Eduardo Perozo and Anthony A Kossiakoff},
doi = {10.1073/pnas.2512532123},
issn = {1091-6490},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {8},
pages = {e2512532123},
abstract = {In prokaryotes, CorA is the primary influx pathway for magnesium, a critical divalent cation in cellular physiology and biochemistry. Mechanistic studies show that homopentameric CorA is regulated through an intracellular [Mg]-dependent negative feedback loop, involving the asymmetric participation of individual subunits. To understand the connection between asymmetry and activation, we used single-particle cryo-EM to solve sixteen structures of nanodisc-reconstituted CorA. We utilized conformation-specific synthetic antibodies to stabilize subtle but significant conformational differences in the cryo-EM structures. Our results demonstrate that CorA exists as a set of conformational ensembles, where population size inversely correlates with intracellular Mg concentration. These ensembles include channels with a variety of pore conformations, both constricted and dilated, suggesting a spectrum of active CorA functional states. The ensembles connect asymmetric structural transitions in the cytoplasmic domain with conformational changes in the permeation pathway via an electrostatic network, ultimately controlling channel-gating events. We believe that these results establish a framework for understanding magnesium homeostasis in prokaryotic systems.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Yao, Zi; Zhao, Fangzhu; Miao, Kun; Peters-Clarke, Trenton M; Zhang, Yun; Ganjave, Snehal D; Vázquez-Maldonado, Angel L; Wu, Yan; Kumru, Kaan; Jumaa, Hammam; Leung, Kevin K; Wells, James A
Targeted shedding of extracellular membrane proteins by induced protease recruitment Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid41756920,
title = {Targeted shedding of extracellular membrane proteins by induced protease recruitment},
author = {Zi Yao and Fangzhu Zhao and Kun Miao and Trenton M Peters-Clarke and Yun Zhang and Snehal D Ganjave and Angel L Vázquez-Maldonado and Yan Wu and Kaan Kumru and Hammam Jumaa and Kevin K Leung and James A Wells},
doi = {10.64898/2026.02.17.706468},
issn = {2692-8205},
year = {2026},
date = {2026-02-01},
urldate = {2026-02-01},
journal = {bioRxiv},
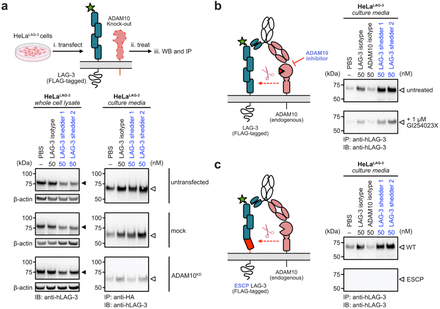
abstract = {Extracellular targeted protein degradation has emerged as a promising therapeutic modality to eliminate proteins of interest (POIs) at the cell surface, by using bifunctional molecules to recruit natural recycling receptors or membrane-bound E3 ligases that redirect POIs to the lysosome. Another natural mechanism involves extracellular proteases that cleave and shed extracellular domains. Here, we exploit this endogenous mechanism by engineering bispecific antibody , that recruit a classic sheddase ADAM10 to POIs, inducing selective ectodomain shedding. We first targeted the immune checkpoint receptor LAG-3 and observed robust depletion of surface LAG-3 accompanied by accumulation of soluble LAG-3 fragments in both engineered cell lines and primary human T cells. Using biochemical and imaging assays, we confirmed that this antibody-induced shedding is restricted to extracellular protease activity and occurs independently of lysosomal trafficking. Notably, induced shedding of LAG-3 on activated primary T cells partially alleviated inhibitory signaling and reinvigorated IFN secretion. We extended the scope of induced shedding by developing that recognize synthetic epitope-tags that enabling rapid assessment of substrate compatibility across diverse targets. Using this platform, we identified multiple immune modulatory cell-surface receptors, including IL6Rα, CD62L and MIC-A that can be targeted for shedding. In summary, this work establishes a new paradigm for targeted extracellular proteolysis and expands the toolkit for studying extracellular proteolysis with potential therapeutic benefit.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Adams, Jarrett J.; Blazer, Levi L.; Chen, Gang; Chung, Jacky; Mallette, Evan; Singer, Alexander U.; Mendiratta, Sanjeev K.; Bandyopadhyay, Sanjay; Singh, Arun K.; Kalita, Pankaj; Karimi, Minoo; Cardarelli, Lia; Geyer, C. Ronald; Michnick, Stephen W.; Sidhu, Sachdev S.
A tetravalent PD-L1/VEGF bispecific antibody blocks tumor growth In Vivo Journal Article
In: Disease and Therapeutics, vol. 2, 2026, ISSN: 3050-9998.
@article{Adams2026,
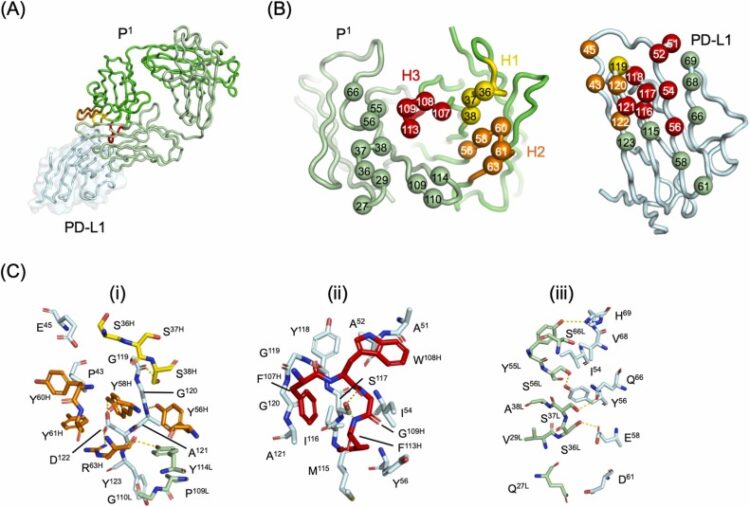
title = {A tetravalent PD-L1/VEGF bispecific antibody blocks tumor growth In Vivo},
author = {Jarrett J. Adams and Levi L. Blazer and Gang Chen and Jacky Chung and Evan Mallette and Alexander U. Singer and Sanjeev K. Mendiratta and Sanjay Bandyopadhyay and Arun K. Singh and Pankaj Kalita and Minoo Karimi and Lia Cardarelli and C. Ronald Geyer and Stephen W. Michnick and Sachdev S. Sidhu},
doi = {10.1016/j.dist.2026.100006},
issn = {3050-9998},
year = {2026},
date = {2026-01-06},
urldate = {2026-00-00},
journal = {Disease and Therapeutics},
volume = {2},
publisher = {Elsevier BV},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Du, Jiale; Andree, Gisele A; Horn-Ghetko, Daniel; Stier, Luca; Singh, Jaspal; Kostrhon, Sebastian; Kiss, Leo; Mann, Matthias; Sidhu, Sachdev S; Schulman, Brenda A
E2 variants for probing E3 ubiquitin ligase activities Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 1, pp. e2524899122, 2026, ISSN: 1091-6490.
@article{pmid41481455,
title = {E2 variants for probing E3 ubiquitin ligase activities},
author = {Jiale Du and Gisele A Andree and Daniel Horn-Ghetko and Luca Stier and Jaspal Singh and Sebastian Kostrhon and Leo Kiss and Matthias Mann and Sachdev S Sidhu and Brenda A Schulman},
doi = {10.1073/pnas.2524899122},
issn = {1091-6490},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {1},
pages = {e2524899122},
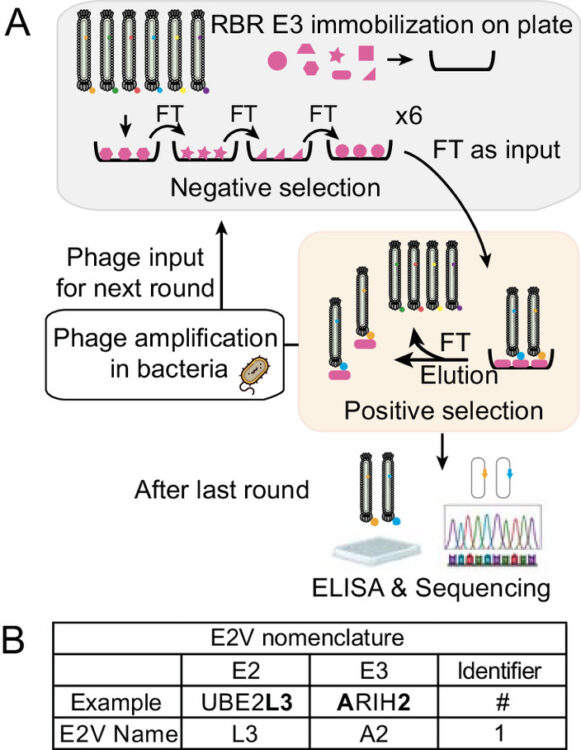
abstract = {E3 ligases partner with E2 enzymes to regulate vast eukaryotic biology. The hierarchical nature of these pairings, with >600 E3s and ~40 E2s in humans, necessitates that E2s cofunction with numerous different E3s. Here, focusing on E3s in the RING-between-RING (RBR) family and their partner UBE2L3 and UBE2D-family E2s, we report an approach to interrogate selected pathways. We screened phage-displayed libraries of structure-based E2 variants (E2Vs) to discover enzymes with enhanced affinity and specificity toward half of all RBR E3 ligases (ARIH1, ARIH2, ANKIB1, CUL9, HOIL1, HOIP, and RNF14). Collectively, these E2Vs allowed distinguishing actions of different cofunctioning E3s, obtaining high-resolution cryogenic Electron Microscopy (cryo-EM) structures of an RBR E3 in the context of a substrate-bound multiprotein complex, and profiling an endogenous RBR E3 response to an extracellular stimulus. Overall, we anticipate that E2V technology will be a generalizable tool to enable in-depth mechanistic and structural analysis of E3 ligase functions, and mapping their activity states and protein partners in cellular signaling cascades.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Kong, Sophie; Peters-Clarke, Trenton M; Delaveris, Corleone S; Phojanakong, Paul; Steri, Veronica; Wells, James A
Cellular consequences, citrullination substrates, and antigenicity resulting from wild-type and targeted PAD4 on cell surfaces Journal Article
In: bioRxiv, 2026, ISSN: 2692-8205.
@article{pmid41542531,
title = {Cellular consequences, citrullination substrates, and antigenicity resulting from wild-type and targeted PAD4 on cell surfaces},
author = {Sophie Kong and Trenton M Peters-Clarke and Corleone S Delaveris and Paul Phojanakong and Veronica Steri and James A Wells},
doi = {10.64898/2026.01.05.696859},
issn = {2692-8205},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {bioRxiv},
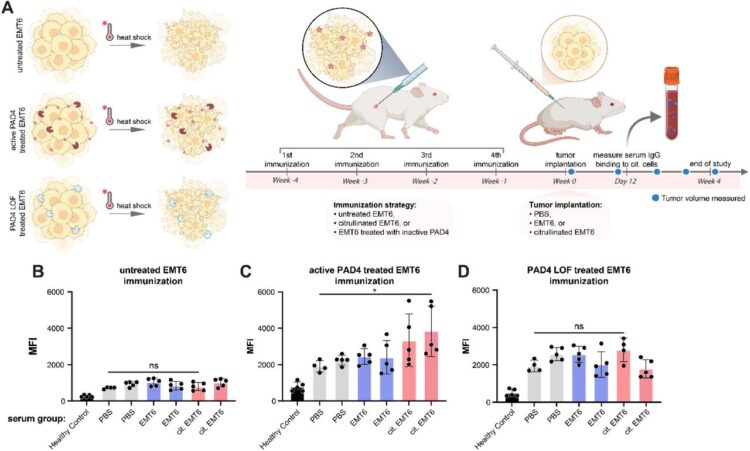
abstract = {Protein arginine deiminase-4 (PAD4) catalyzes hydrolysis of arginine to citrulline in proteins that promotes widespread changes in cellular phenotypes through transcriptional regulation that can induce innate immunity and promote cancer. Overexpression and hyperactivity of PAD4 leads to a form of cell death called NETosis that releases PAD4 to the extracellular space. In excess, release of PAD4 is believed to be a major cause of various autoimmune diseases through the generation of anti-citrulline protein antibodies (ACPAs). Little is known about the specific protein substrates that become citrullinated and lead to autoimmunity, but there is growing evidence that PAD4 can be localized to the cell surface in response to inflammation. Here, we further characterize the cellular consequences for exogenous treatment with PAD4 showing that it induces morphological changes that increase cell migration, a hallmark of cancer. We then devised a more simplified and robust proteomics approach to identify PAD4 substrates. We identified some 1000 endogenously citrullinated peptides from 500 proteins, and 3000 citrullinated peptides from 1300 proteins upon exogenous addition of PAD4 both inside and outside of cells. This extracellular set can be further augmented by targeting PAD4 to a cancer target, HER2, using a binding protein conjugate. Finally, we studied how citrullinated cells can induce a robust humoral response in a syngeneic vaccine model to produce ACPAs. We believe these studies further our understanding of cell phenotypic consequences of extracellular PAD4 and new PAD4 substrates both inside and outside of cells that are potential neoepitopes for generation of ACPAs.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Kumru, Kaan; Yao, Zi; Holmes, Brandon B; Zhao, Fangzhu; Zhang, Yun; Ferrara, Emilio; Peters-Clarke, Trenton M; Leung, Kevin K; Wells, James A
A cytokine receptor-targeting chimera toolbox for expanding extracellular targeted protein degradation Journal Article
In: Proc Natl Acad Sci U S A, vol. 123, no. 4, pp. e2524129123, 2026, ISSN: 1091-6490.
@article{pmid41564137,
title = {A cytokine receptor-targeting chimera toolbox for expanding extracellular targeted protein degradation},
author = {Kaan Kumru and Zi Yao and Brandon B Holmes and Fangzhu Zhao and Yun Zhang and Emilio Ferrara and Trenton M Peters-Clarke and Kevin K Leung and James A Wells},
doi = {10.1073/pnas.2524129123},
issn = {1091-6490},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Proc Natl Acad Sci U S A},
volume = {123},
number = {4},
pages = {e2524129123},
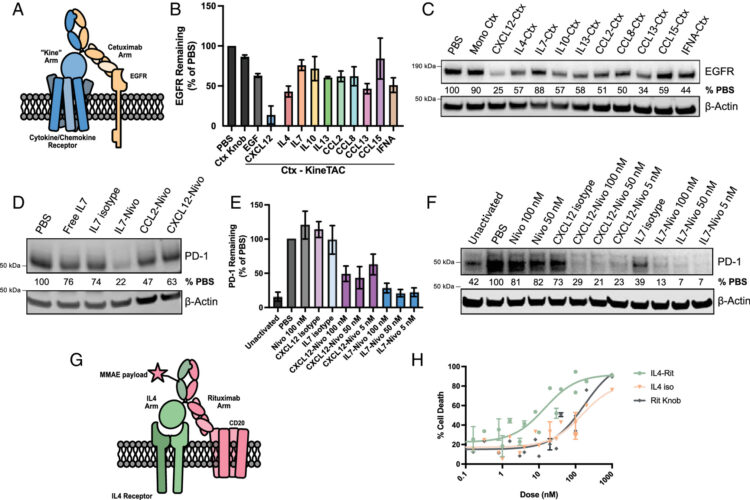
abstract = {Extracellular targeted protein degradation (eTPD) is an important new modality for manipulating the extracellular proteome. However, most eTPD receptors are expressed broadly or are restricted to the liver, limiting specific degradation in other tissues. Cytokine receptor-targeting chimeras (kineTACs) are genetically encoded bispecifics for eTPD that fuse a natural ligand like CXCL12 to an antibody, directing soluble or membrane proteins for lysosomal degradation using the widely expressed chemokine receptor CXCR7 (K. Pance , , 273-281 (2023)]. Here, we dramatically expand the kineTAC toolbox by constructing 81 different kineTACs based on an unbiased list of cytokines, chemokines, and growth factors. Remarkably, 55 of these expressed at suitable levels for analysis without any optimization. Many of these kineTACs bind receptors that have unique cell-type expression profiles, allowing for eTPD in specific cells and tissues, and some were more potent than the original CXCL12-based kineTAC against specific targets. We further show the internalizing capability of a kineTAC can enhance the performance of antibody drug conjugates. We believe these simple, genetically encoded tools will be useful for expanding the applications for optimized or cell type-selective eTPD.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Montaño, José L; Tejus, Vishnu R; Xu, Vee; Condon, Andrew C; Ecker, Andrew K; Langen, Andreas; Leung, Kevin K; Zaro, Balyn W
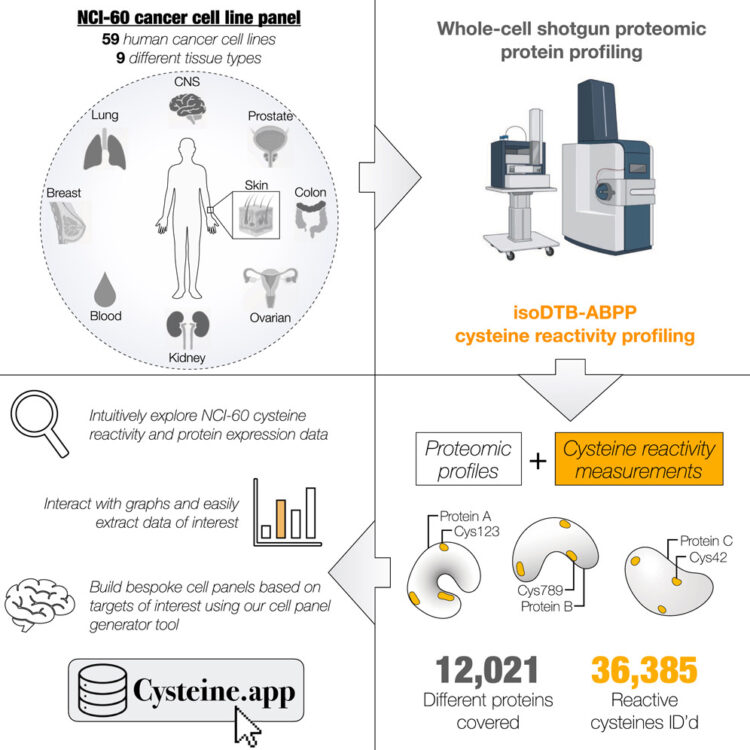
An interactive resource mapping the proteome and reactive cysteine landscape across the NCI-60 reveals cell and tissue-specific profiles Journal Article
In: Cell Chem Biol, vol. 33, no. 1, pp. 117–131.e5, 2026, ISSN: 2451-9448.
@article{pmid41478280,
title = {An interactive resource mapping the proteome and reactive cysteine landscape across the NCI-60 reveals cell and tissue-specific profiles},
author = {José L Montaño and Vishnu R Tejus and Vee Xu and Andrew C Condon and Andrew K Ecker and Andreas Langen and Kevin K Leung and Balyn W Zaro},
doi = {10.1016/j.chembiol.2025.12.003},
issn = {2451-9448},
year = {2026},
date = {2026-01-01},
urldate = {2026-01-01},
journal = {Cell Chem Biol},
volume = {33},
number = {1},
pages = {117--131.e5},
abstract = {The NCI-60 cancer cell line panel is one of the most extensively characterized and publicly accessible resources in cancer research. Here, we expand this platform by generating a comprehensive proteomic and cysteine reactivity atlas using shotgun proteomics and quantitative chemoproteomics. We quantified over 12,000 proteins and identified more than 36,000 reactive cysteines, including over 10,000 hyperreactive sites, across the panel. Our analyses reveal widespread heterogeneity in cysteine reactivity, while conserved hyperreactive cysteines strongly correlate with functional roles. Tissue-specific cysteine reactivity occurred independently of protein abundance, highlighting context-dependent regulation. To enable exploration and integration with existing datasets, we developed an interactive online database that allows users to visualize protein and cysteine coverage and design custom cell line panels. Together, these data and tools illuminate the landscape of cysteine reactivity in cancer and provide a foundational resource to advance future studies in protein function, redox biology, and covalent drug discovery.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}