Welcome to the Recombinant Antibody Network
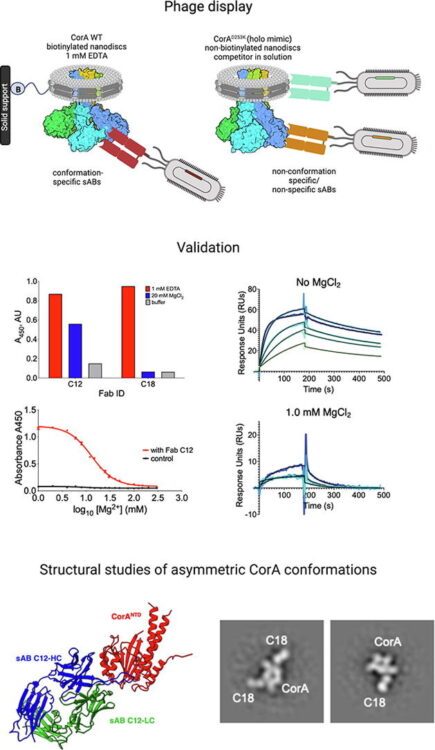
Latest Publications
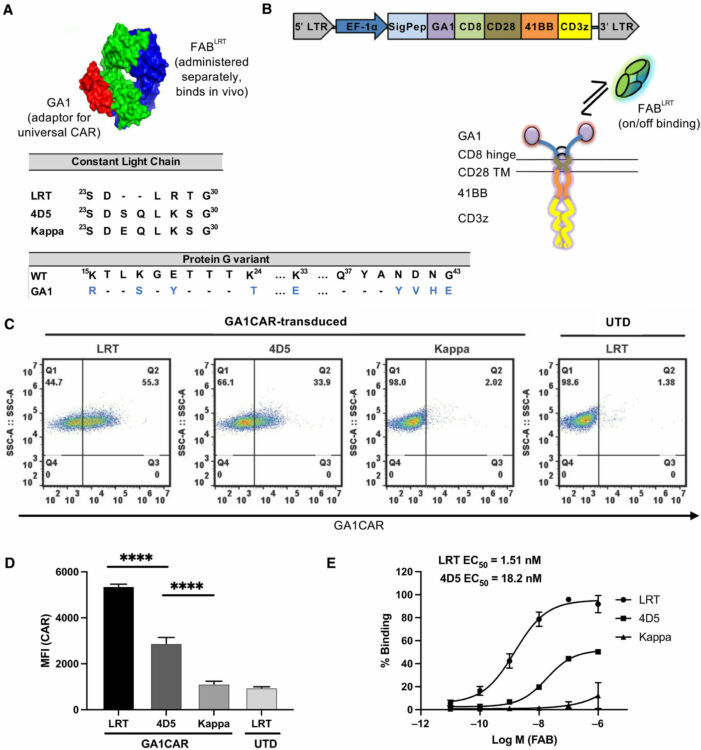
A universal chimeric antigen receptor (CAR)-fragment antibody binder (FAB) split system for cancer immunotherapy Journal Article
In: Sci Adv, vol. 11, no. 27, pp. eadv4937, 2025, ISSN: 2375-2548.
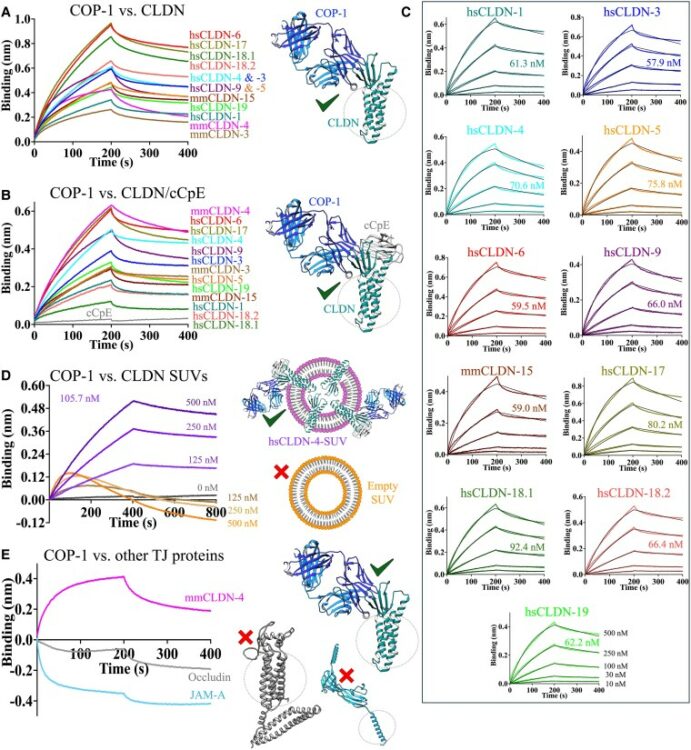
Biophysical basis of tight junction barrier modulation by a pan-claudin-binding molecule Journal Article
In: PNAS Nexus, vol. 4, no. 6, pp. pgaf189, 2025, ISSN: 2752-6542.
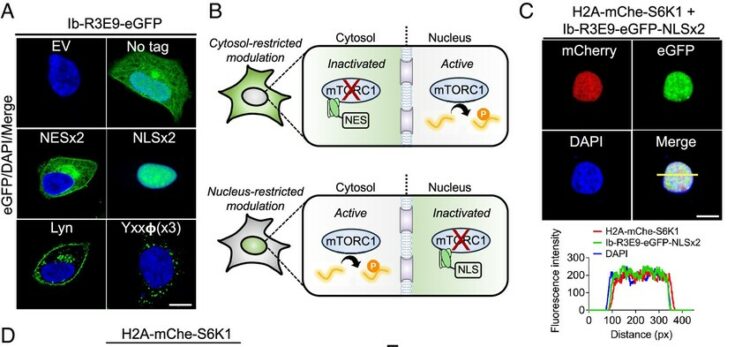
Conformation-specific synthetic intrabodies modulate mTOR signaling with subcellular spatial resolution Journal Article
In: Proc Natl Acad Sci U S A, vol. 122, no. 24, pp. e2424679122, 2025, ISSN: 1091-6490.
Latest News
Recombinant Antibody Network Partners with Bristol Myers Squibb to Develop Novel Therapies
The Recombinant Antibody Network (RAN), a consortium comprising research groups from UC San Francisco, the…
Absolute Antibody Partners with the Recombinant Antibody Network to Facilitate Access to Engineered Recombinant Antibodies
Absolute Antibody Ltd., an industry-leading provider of recombinant antibody products and services, has announced a…
RAN to collaborate with Celgene on cancer therapeutics development
The RAN has recently signed a 3-year $25M agreement with the Celgene Corporation to develop next-generation,…
